Time series data is a sequence of data points indexed by time, where each data point is recorded at a specific interval. TimesFM is a foundation model pretrained on a large corpus of time series data. Its architecture is a decoder-only transformer, similar to LLMs, which learns to predict the next part of a time series from previous data. We can use the follow pipeline to detect anomalies in time series data and periodically learn from incoming data to improve our timesfm predictions.
pip install timesfm[torch]pip install 'apache_beam[interactive,gcp,test] == 2.67.0'pip install google-generativeai
Ordered Sliding Window
import logging
import apache_beam as beam
from apache_beam.coders import BooleanCoder
from apache_beam.coders import PickleCoder
from apache_beam.coders import TimestampCoder
from apache_beam.transforms.timeutil import TimeDomain
from apache_beam.transforms.userstate import OrderedListStateSpec
from apache_beam.transforms.userstate import ReadModifyWriteStateSpec
from apache_beam.transforms.userstate import TimerSpec
from apache_beam.transforms.userstate import on_timer
from apache_beam.utils.timestamp import MAX_TIMESTAMP
from apache_beam.utils.timestamp import Timestamp
_LOGGER = logging.getLogger(__name__)
logging.basicConfig(level=logging.INFO)
_LOGGER.setLevel(logging.INFO)
class OrderedSlidingWindowFn(beam.DoFn):
ORDERED_BUFFER_STATE = OrderedListStateSpec('ordered_buffer', PickleCoder())
WINDOW_TIMER = TimerSpec('window_timer', TimeDomain.WATERMARK)
TIMER_STATE = ReadModifyWriteStateSpec('timer_state', BooleanCoder())
EARLIEST_TS_STATE = ReadModifyWriteStateSpec('earliest_ts', TimestampCoder())
def __init__(self, window_size, slide_interval):
self.window_size = window_size
self.slide_interval = slide_interval
def start_bundle(self):
_LOGGER.debug("start bundle")
def finish_bundle(self):
_LOGGER.debug("finish bundle")
def process(
self,
element,
timestamp=beam.DoFn.TimestampParam,
ordered_buffer=beam.DoFn.StateParam(ORDERED_BUFFER_STATE),
window_timer=beam.DoFn.TimerParam(WINDOW_TIMER),
timer_state=beam.DoFn.StateParam(TIMER_STATE),
earliest_ts_state=beam.DoFn.StateParam(EARLIEST_TS_STATE)):
_, value = element
ordered_buffer.add((timestamp, value))
_LOGGER.debug("receive %s at %s", element, timestamp)
timer_started = timer_state.read()
earliest = earliest_ts_state.read()
if not earliest or earliest > timestamp:
earliest_ts_state.write(timestamp)
if not timer_started:
earliest_ts_state.write(timestamp)
first_slide_start = int(
timestamp.micros / 1e6 // self.slide_interval) * self.slide_interval
first_slide_start_ts = Timestamp.of(first_slide_start)
first_window_end_ts = first_slide_start_ts + self.window_size
_LOGGER.debug("set timer to %s", first_window_end_ts)
window_timer.set(first_window_end_ts)
timer_state.write(True)
return []
@on_timer(WINDOW_TIMER)
def on_timer(
self,
key=beam.DoFn.KeyParam,
fire_ts=beam.DoFn.TimestampParam,
ordered_buffer=beam.DoFn.StateParam(ORDERED_BUFFER_STATE),
window_timer=beam.DoFn.TimerParam(WINDOW_TIMER),
timer_state=beam.DoFn.StateParam(TIMER_STATE),
earliest_ts_state=beam.DoFn.StateParam(EARLIEST_TS_STATE)):
_LOGGER.debug("timer fire at %s", fire_ts)
window_end_ts = fire_ts
window_start_ts = window_end_ts - self.window_size
window_values = list(
ordered_buffer.read_range(window_start_ts, window_end_ts))
_LOGGER.debug(
"window start: %s, window end: %s", window_start_ts, window_end_ts)
_LOGGER.debug("windowed data in buffer %s", str(window_values))
if window_values:
yield (key, (window_start_ts, window_end_ts, window_values))
next_window_end_ts = fire_ts + self.slide_interval
next_window_start_ts = window_start_ts + self.slide_interval
earliest_ts = earliest_ts_state.read()
ordered_buffer.clear_range(earliest_ts, next_window_start_ts)
remaining_data = list(
ordered_buffer.read_range(next_window_start_ts, MAX_TIMESTAMP))
if not remaining_data:
timer_state.clear()
earliest_ts_state.write(next_window_start_ts)
return
_LOGGER.debug("set timer to %s", next_window_end_ts)
window_timer.set(next_window_end_ts)
class FillGapsFn(beam.DoFn):
def __init__(self, expected_interval: float):
"""
Args:
expected_interval: The expected time delta between elements, in seconds.
"""
self.expected_interval = expected_interval
def process(self, element):
key, (window_start_ts, window_end_ts, window_elements) = element
received_data = {
round(float(ts.micros / 1e6), 5): val
for ts, val in window_elements
}
start_sec = float(window_start_ts.micros / 1e6)
end_sec = float(window_end_ts.micros / 1e6)
filled_values = []
current_ts_sec = start_sec
while current_ts_sec < end_sec:
lookup_ts = round(current_ts_sec, 5)
if lookup_ts in received_data:
filled_values.append(float(received_data[lookup_ts]))
else:
filled_values.append('NaN')
current_ts_sec += self.expected_interval
yield (key, (window_start_ts, window_end_ts, filled_values))
Model Handler
import apache_beam as beam
from apache_beam.ml.inference.base import ModelHandler
import timesfm
import logging
import numpy as np
import os
from google.cloud import storage
from apache_beam.io.gcp.gcsio import GcsIO
from apache_beam.utils.timestamp import Timestamp
class LatestModelCheckpointLoader(beam.PTransform):
"""A PTransform that finds the latest model checkpoint in a GCS path."""
def __init__(self, gcs_bucket, gcs_prefix):
self.gcs_bucket = gcs_bucket
self.gcs_prefix = gcs_prefix
def expand(self, pcoll):
return pcoll | "FindLatestModel" >> beam.Map(self._find_latest_model_path)
def _find_latest_model_path(self, _):
try:
storage_client = storage.Client()
blobs = storage_client.list_blobs(self.gcs_bucket, prefix=self.gcs_prefix)
# Filter for model files and find the most recent one
model_blobs = [b for b in blobs if b.name.endswith(".pth")]
latest_blob = max(model_blobs, key=lambda b: b.time_created, default=None)
if latest_blob:
path = f"gs://{self.gcs_bucket}/{latest_blob.name}"
logging.info(f"Found latest finetuned model at: {path}")
return path
except Exception as e:
logging.error(f"Error finding latest model in GCS: {e}")
# Return a path to the base model if no finetuned one exists or an error occurs
base_model = "google/timesfm-1.0-200m-pytorch"
logging.info(f"No finetuned model found. Using base model: {base_model}")
return base_model
class DynamicTimesFmModelHandler(ModelHandler[np.ndarray, np.ndarray, timesfm.TimesFm]):
"""
A model handler that loads a TimesFM model from a dynamic path (GCS or Hugging Face).
The model path is provided as a side input to RunInference.
"""
def __init__(self, model_uri: str, hparams):
self._hparams = hparams
self._model = None
self._model_uri = model_uri
self._context_len = hparams.context_len
self._horizon_len = hparams.horizon_len
def load_model(self) -> timesfm.TimesFm:
"""Loads a model from the handler's current model_uri."""
logging.info(f"Loading TimesFM model from path: {self._model_uri}...")
checkpoint_config = {}
if self._model_uri.startswith("gs://"):
try:
gcs = GcsIO()
file_name = os.path.basename(self._model_uri)
local_path = f"/tmp/{file_name}"
with gcs.open(self._model_uri, 'rb') as f_in, open(local_path, 'wb') as f_out:
f_out.write(f_in.read())
checkpoint_config['path'] = local_path
logging.info(f"Downloaded model from GCS to {local_path}")
except Exception as e:
logging.error(f"Failed to download model from GCS: {e}. Check path and permissions.")
raise e # Re-raise the exception to fail fast if the model can't be loaded.
else:
checkpoint_config['huggingface_repo_id'] = self._model_uri
self._model = timesfm.TimesFm(
hparams=self._hparams,
checkpoint=timesfm.TimesFmCheckpoint(**checkpoint_config)
)
logging.info("TimesFM model loaded successfully.")
return self._model
def update_model_path(self, model_path: str):
"""
This method is called by RunInference when a new model metadata is available
from the side input. It updates the model URI that `load_model` will use.
"""
if not model_path:
logging.info("Received an empty model path update. No action taken.")
return
logging.info(f"Received model update. New model URI: {model_path}")
self._model_uri = model_path
self._model = self.load_model()
logging.info("Model has been updated in the handler.")
def run_inference(self, batch, model, inference_args=None):
"""
Runs inference on a batch of data.
Note: While this is a standard method for ModelHandler, we will call the
model's `forecast` method directly in our DoFn for clarity.
"""
# print("Running inference on batch:", batch)
# logging.info(f"Running inference on batch:", batch)
anomalies_found = []
key, (window_start_ts, _, values_array) = batch[0]
# A window must have enough data for both context and horizon.
# if len(values_array) < self.context_len + self.horizon_len:
# return
current_context = np.array(values_array[:self._context_len])
actual_horizon_values = np.array(
values_array[self._context_len:self._context_len + self._horizon_len])
print("Current context shape:", current_context.shape)
print("Actual horizon values shape:", actual_horizon_values.shape)
point_forecast, experimental_quantile_forecast = model.forecast(
[current_context],
freq=[0],
)
current_predicted_horizon_values = point_forecast[
0, :, 0] if point_forecast.ndim == 3 else point_forecast[0]
current_q20_values = experimental_quantile_forecast[0, :, 2]
current_q30_values = experimental_quantile_forecast[0, :, 3]
current_q70_values = experimental_quantile_forecast[0, :, 7]
current_q80_values = experimental_quantile_forecast[0, :, 8]
for j in range(len(actual_horizon_values)):
current_actual = actual_horizon_values[j]
point_Q1 = np.nanmean([current_q20_values[j], current_q30_values[j]])
point_Q3 = np.nanmean([current_q70_values[j], current_q80_values[j]])
point_IQR = point_Q3 - point_Q1
upper_thresh = point_Q3 + 1.5 * point_IQR
lower_thresh = point_Q1 - 1.5 * point_IQR
if current_actual > upper_thresh or current_actual < lower_thresh:
score = (current_actual - upper_thresh
) / point_IQR if current_actual > upper_thresh else (
lower_thresh - current_actual) / point_IQR
anomaly_timestamp_seconds = (window_start_ts.micros / 1e6) + (
self._context_len + j)
index_in_window = self._context_len + j
anomalies_found.append({
'key': key,
'timestamp': Timestamp(anomaly_timestamp_seconds),
'index_in_window': index_in_window,
'actual_value': current_actual,
'predicted_value': current_predicted_horizon_values[j],
'is_anomaly': True,
'outlier_score': score,
'lower_bound': lower_thresh,
'upper_bound': upper_thresh,
})
payload = {
"start_ts_micros": window_start_ts.micros,
"predicted_values": current_predicted_horizon_values.tolist(),
"q20_values": current_q20_values.tolist(),
"q30_values": current_q30_values.tolist(),
"q70_values": current_q70_values.tolist(),
"q80_values": current_q80_values.tolist(),
"anomalies": anomalies_found, # Your original list is now inside the dictionary
"actual_horizon_values": actual_horizon_values.tolist()
}
result_with_context = (batch[0], payload)
return [result_with_context]
See https://github.com/google-research/timesfm/blob/master/README.md for updated APIs. Loaded PyTorch TimesFM, likely because python version is 3.11.13 (main, Jun 4 2025, 08:57:29) [GCC 11.4.0].
LLM Classifier
import apache_beam as beam
import google.generativeai as genai
import logging
import os
import re
import json
import numpy as np
from apache_beam.utils.timestamp import Timestamp
from dotenv import load_dotenv
from apache_beam.transforms.userstate import BagStateSpec
import apache_beam as beam
import json
import numpy as np
from apache_beam.coders.coders import PickleCoder
from apache_beam.transforms.userstate import BagStateSpec, ReadModifyWriteStateSpec, TimerSpec, on_timer
class CustomJsonEncoderForLLM(json.JSONEncoder):
"""Encodes special types like Timestamp and numpy objects into JSON."""
def default(self, obj):
if isinstance(obj, Timestamp):
# Store as a dict with a special key for easy decoding
return {'__timestamp__': True, 'micros': obj.micros}
if isinstance(obj, np.integer):
return int(obj)
if isinstance(obj, np.floating):
return float(obj)
if isinstance(obj, np.ndarray):
return obj.tolist()
return super().default(obj)
def custom_json_decoder(dct):
"""Decodes a Timestamp object from our custom dict format."""
if '__timestamp__' in dct:
return Timestamp(micros=dct['micros'])
return dct
class JsonCoderWithNumpyAndTimestamp(beam.coders.Coder):
"""A custom Beam Coder that handles JSON serialization for Timestamps and numpy types."""
def encode(self, value):
return json.dumps(value, cls=CustomJsonEncoderForLLM).encode('utf-8')
def decode(self, encoded):
return json.loads(encoded.decode('utf-8'), object_hook=custom_json_decoder)
def is_deterministic(self):
return True
# It's highly recommended to manage API keys via GCP Secret Manager
# and access them as environment variables in your Dataflow job.
# genai.configure(api_key=os.environ["GEMINI_API_KEY"])
class LLMClassifierFn(beam.DoFn):
"""
Takes an anomaly, formats a detailed prompt with surrounding context,
calls the Gemini model to classify it, and routes the original data
based on the model's decision.
This DoFn is stateful, deferring anomalies that occur too close to
the end of a window until a subsequent window provides enough context.
"""
DEFERRED_ANOMALIES_STATE = BagStateSpec(
'deferred_anomalies', coder=JsonCoderWithNumpyAndTimestamp())
YIELD_BUFFER_STATE = ReadModifyWriteStateSpec('yield_buffer', PickleCoder())
# <<< CHANGE: Define a timer and a state to track if it's set
EXPIRY_TIMER = TimerSpec('expiry', beam.TimeDomain.WATERMARK)
# <<< CHANGE: Add state to track the last yielded timestamp
LAST_YIELDED_TIMESTAMP_STATE = ReadModifyWriteStateSpec('last_yielded_ts', PickleCoder())
def __init__(self, secret, context_points=25, slide_interval=128, expected_interval_secs=1):
self.context_points = context_points
self._model = None
self.secret = secret
self.slide_interval = slide_interval
self.expected_interval_micros = expected_interval_secs * 1_000_000
self._last_window_data = None
def setup(self):
# Configure the generative model
genai.configure(api_key=self.secret)
logging.getLogger().setLevel(logging.INFO)
generation_config = {
"temperature": 0.2,
"top_p": 1,
"top_k": 1,
"max_output_tokens": 256,
"response_mime_type": "application/json",
}
# For a full list of safety settings, see the Gemini API documentation
safety_settings = [
{"category": "HARM_CATEGORY_HARASSMENT", "threshold": "BLOCK_NONE"},
{"category": "HARM_CATEGORY_HATE_SPEECH", "threshold": "BLOCK_NONE"},
]
self._model = genai.GenerativeModel(
model_name="gemini-1.5-flash-latest",
generation_config=generation_config,
safety_settings=safety_settings
)
logging.info("Gemini Model has been successfully initialized.")
def _build_prompt(self, anomaly_data, context_before, context_after):
mean_before = np.mean(context_before) if context_before.size > 0 else 0
mean_after = np.mean(context_after) if context_after.size > 0 else 0
std_before = np.std(context_before) if context_before.size > 0 else 0
std_after = np.std(context_after) if context_after.size > 0 else 0
return f"""
You are an expert time-series analyst classifying an outlier from NYC taxi pickup data.
Normal behavior includes daily and weekly cyclical patterns.
**1. Outlier Context:**
* **--> The Outlier:**
* **Timestamp:** {Timestamp(micros=anomaly_data['timestamp'].micros)}
* **Actual Value:** {anomaly_data['actual_value']:.2f}
* **Predicted Value:** {anomaly_data['predicted_value']:.2f}
* **Anomaly Upper Bound:** {anomaly_data['upper_bound']:.2f}
* **Anomaly Lower Bound:** {anomaly_data['lower_bound']:.2f}
**2. Data Surrounding the Outlier:**
* **Data Before ({len(context_before)} points):** {np.round(context_before, 2).tolist()}
* **Data After ({len(context_after)} points):** {np.round(context_after, 2).tolist()}
**3. Statistical Context:**
* **Mean Before:** {mean_before:.2f}
* **Mean After:** {mean_after:.2f}
* **Std. Dev. Before:** {std_before:.2f}
* **Std. Dev. After:** {std_after:.2f}
**4. Your Task:**
**Step 1: Analyze the Evidence.** In a few sentences, describe the behavior of the data *after* the outlier. Does it quickly revert to the "Predicted Value" or the "Mean Before"? Or does it establish a new level, closer to the "Mean After"?
**Step 2: Make a Decision.** Classify the outlier.
* **REMOVE:** If it's a transient, one-off event. This is likely if the data after the outlier rapidly returns to the established pattern.
* **KEEP:** If it signifies a sustained shift in the pattern that the model should learn from. This is likely if the `Mean After` has shifted significantly.
**Step 3: Provide Final Output.** Respond with a single JSON object. Do not add any text outside the JSON block.
{ {
"reasoning_steps": "Your analysis from Step 1 goes here.",
"decision": "KEEP or REMOVE",
"confidence_score": <A float between 0.0 and 1.0>
} }
"""
def process(self, element,
deferred_anomalies=beam.DoFn.StateParam(DEFERRED_ANOMALIES_STATE),
yield_buffer=beam.DoFn.StateParam(YIELD_BUFFER_STATE),
expiry_timer=beam.DoFn.TimerParam(EXPIRY_TIMER)):
key, data = element
window_start_ts = data['window_start_ts']
# Set a timer to fire based on the event time of the current element.
# Each new element will push the timer forward. The timer will only
# fire when a gap in the input stream occurs, allowing the buffer
# to contain data from multiple consecutive sliding windows.
# We set it far enough ahead to allow the next window's data to arrive.
grace_period_secs = self.slide_interval * 2
expiry_timer.set(window_start_ts + grace_period_secs)
anomalies_in_window = data.get('anomalies', [])
values_in_element = data.get('values_array', [])
for anomaly in anomalies_in_window:
deferred_anomalies.add(anomaly)
buffer = yield_buffer.read() or {}
for i, value in enumerate(values_in_element):
point_timestamp = Timestamp(micros=window_start_ts.micros + (i * self.expected_interval_micros))
buffer[point_timestamp] = value
yield_buffer.write(buffer)
@on_timer(EXPIRY_TIMER)
def on_expiry_timer(
self,
deferred_anomalies=beam.DoFn.StateParam(DEFERRED_ANOMALIES_STATE),
yield_buffer=beam.DoFn.StateParam(YIELD_BUFFER_STATE),
# <<< CHANGE: Add the new state parameter here
last_yielded_ts_state=beam.DoFn.StateParam(LAST_YIELDED_TIMESTAMP_STATE)):
all_anomalies_to_consider = list(deferred_anomalies.read())
buffered_points_map = yield_buffer.read() or {}
if not buffered_points_map:
return
sorted_points = sorted(buffered_points_map.items())
all_timestamps = [ts for ts, val in sorted_points]
all_values = [val for ts, val in sorted_points]
anomalies_to_process_now = []
prompts_to_batch = []
final_deferred = []
for anomaly_data in all_anomalies_to_consider:
anomaly_ts = anomaly_data['timestamp']
try:
idx_in_full_data = all_timestamps.index(anomaly_ts)
if (idx_in_full_data + self.context_points) < len(all_values):
start_ctx = max(0, idx_in_full_data - self.context_points)
end_ctx = idx_in_full_data + self.context_points + 1
context_before = np.array(all_values[start_ctx:idx_in_full_data])
context_after = np.array(all_values[idx_in_full_data + 1:end_ctx])
anomaly_data['index_in_window'] = idx_in_full_data
prompt = self._build_prompt(anomaly_data, context_before, context_after)
prompts_to_batch.append(prompt)
anomalies_to_process_now.append(anomaly_data)
else:
final_deferred.append(anomaly_data)
except ValueError:
final_deferred.append(anomaly_data)
if prompts_to_batch:
try:
logging.info(f"Sending a batch of {len(prompts_to_batch)} prompts to the LLM.")
responses = self._model.generate_content(prompts_to_batch)
for anomaly_data, response in zip(anomalies_to_process_now, responses):
try:
response_data = json.loads(response.text)
decision = response_data.get('decision', 'KEEP').strip().upper()
idx = anomaly_data['index_in_window']
if decision == 'REMOVE':
logging.warning(f"LLM decided to REMOVE anomaly at {anomaly_data['timestamp']}. Imputing value.")
all_values[idx] = anomaly_data['predicted_value']
except (json.JSONDecodeError, AttributeError) as e:
logging.error(f"Error processing LLM response for {anomaly_data['timestamp']}: {e}. Defaulting to KEEP.")
except Exception as e:
logging.error(f"Error calling LLM with a batch: {e}. Defaulting to KEEP for all.")
# <<< CHANGE: New logic to yield only new data
last_yielded_ts = last_yielded_ts_state.read()
latest_ts_in_batch = None
for i, (ts, original_val) in enumerate(sorted_points):
# Only yield points that are newer than the last batch we yielded
if last_yielded_ts is None or ts > last_yielded_ts:
yield {
'timestamp': ts,
'value': all_values[i]
}
latest_ts_in_batch = ts
# After yielding, update the state with the latest timestamp from this batch
if latest_ts_in_batch:
last_yielded_ts_state.write(latest_ts_in_batch)
# Prune the buffer. We need to keep enough historical data to serve
# as `context_before` for the anomalies that we are re-deferring.
if latest_ts_in_batch:
all_buffered_points = yield_buffer.read() or {}
# Find the earliest timestamp we need to keep. This will be
# `context_points` before the last yielded point, ensuring
# context is available for the next batch.
try:
last_yielded_index = all_timestamps.index(latest_ts_in_batch)
context_start_index = max(0, last_yielded_index - self.context_points)
context_start_ts = all_timestamps[context_start_index]
pruned_buffer = {
ts: val
for ts, val in all_buffered_points.items()
if ts >= context_start_ts
}
yield_buffer.write(pruned_buffer)
except ValueError:
# This can happen if the buffer is in an inconsistent state.
# As a fallback, we clear it if we aren't deferring anything.
logging.warning(
f"Could not find last yielded timestamp "
f"{latest_ts_in_batch} in buffer for pruning."
)
if not final_deferred:
yield_buffer.clear()
elif not final_deferred:
# If we didn't yield anything and we're not deferring anything,
# the buffer is fully processed and can be cleared.
yield_buffer.clear()
# Re-add anomalies that couldn't be processed to the state so they can
# be considered in the next firing.
deferred_anomalies.clear()
if final_deferred:
logging.info(f"Re-deferring {len(final_deferred)} anomalies due to insufficient context.")
for anomaly in final_deferred:
deferred_anomalies.add(anomaly)
Finetuning Component
"""
TimesFM Finetuner: A flexible framework for finetuning TimesFM models on custom datasets.
"""
import logging
import os
from abc import ABC, abstractmethod
from dataclasses import dataclass, field
from typing import Any, Callable, Dict, List, Optional
import torch
import torch.distributed as dist
import torch.nn as nn
from torch.nn.parallel import DistributedDataParallel as DDP
from torch.utils.data import DataLoader, Dataset
from timesfm.pytorch_patched_decoder import create_quantiles
import wandb
class MetricsLogger(ABC):
"""Abstract base class for logging metrics during training.
This class defines the interface for logging metrics during model training.
Concrete implementations can log to different backends (e.g., WandB, TensorBoard).
"""
@abstractmethod
def log_metrics(self,
metrics: Dict[str, Any],
step: Optional[int] = None) -> None:
"""Log metrics to the specified backend.
Args:
metrics: Dictionary containing metric names and values.
step: Optional step number or epoch for the metrics.
"""
pass
@abstractmethod
def close(self) -> None:
"""Clean up any resources used by the logger."""
pass
class WandBLogger(MetricsLogger):
"""Weights & Biases implementation of metrics logging.
Args:
project: Name of the W&B project.
config: Configuration dictionary to log.
rank: Process rank in distributed training.
"""
def __init__(self, project: str, config: Dict[str, Any], rank: int = 0):
self.rank = rank
if rank == 0:
wandb.init(project=project, config=config)
def log_metrics(self,
metrics: Dict[str, Any],
step: Optional[int] = None) -> None:
"""Log metrics to W&B if on the main process.
Args:
metrics: Dictionary of metrics to log.
step: Current training step or epoch.
"""
if self.rank == 0:
wandb.log(metrics, step=step)
def close(self) -> None:
"""Finish the W&B run if on the main process."""
if self.rank == 0:
wandb.finish()
class DistributedManager:
"""Manages distributed training setup and cleanup.
Args:
world_size: Total number of processes.
rank: Process rank.
master_addr: Address of the master process.
master_port: Port for distributed communication.
backend: PyTorch distributed backend to use.
"""
def __init__(
self,
world_size: int,
rank: int,
master_addr: str = "localhost",
master_port: str = "12358",
backend: str = "nccl",
):
self.world_size = world_size
self.rank = rank
self.master_addr = master_addr
self.master_port = master_port
self.backend = backend
def setup(self) -> None:
"""Initialize the distributed environment."""
os.environ["MASTER_ADDR"] = self.master_addr
os.environ["MASTER_PORT"] = self.master_port
if not dist.is_initialized():
dist.init_process_group(backend=self.backend,
world_size=self.world_size,
rank=self.rank)
def cleanup(self) -> None:
"""Clean up the distributed environment."""
if dist.is_initialized():
dist.destroy_process_group()
@dataclass
class FinetuningConfig:
"""Configuration for model training.
Args:
batch_size: Number of samples per batch.
num_epochs: Number of training epochs.
learning_rate: Initial learning rate.
weight_decay: L2 regularization factor.
freq_type: Frequency, can be [0, 1, 2].
use_quantile_loss: bool = False # Flag to enable/disable quantile loss
quantiles: Optional[List[float]] = None
device: Device to train on ('cuda' or 'cpu').
distributed: Whether to use distributed training.
gpu_ids: List of GPU IDs to use.
master_port: Port for distributed training.
master_addr: Address for distributed training.
use_wandb: Whether to use Weights & Biases logging.
wandb_project: W&B project name.
log_every_n_steps: Log metrics every N steps (batches), this is inspired from Pytorch Lightning
val_check_interval: How often within one training epoch to check val metrics. (also from Pytorch Lightning)
Can be: float (0.0-1.0): fraction of epoch (e.g., 0.5 = validate twice per epoch)
int: validate every N batches
"""
batch_size: int = 32
num_epochs: int = 20
learning_rate: float = 1e-4
weight_decay: float = 0.01
freq_type: int = 0
use_quantile_loss: bool = False
quantiles: Optional[List[float]] = None
device: str = "cuda" if torch.cuda.is_available() else "cpu"
distributed: bool = False
gpu_ids: List[int] = field(default_factory=lambda: [0])
master_port: str = "12358"
master_addr: str = "localhost"
use_wandb: bool = False
wandb_project: str = "timesfm-finetuning"
log_every_n_steps: int = 50
val_check_interval: float = 0.5
class TimesFMFinetuner:
"""Handles model training and validation.
Args:
model: PyTorch model to train.
config: Training configuration.
rank: Process rank for distributed training.
loss_fn: Loss function (defaults to MSE).
logger: Optional logging.Logger instance.
"""
def __init__(
self,
model: nn.Module,
config: FinetuningConfig,
rank: int = 0,
loss_fn: Optional[Callable] = None,
logger: Optional[logging.Logger] = None,
):
self.model = model
self.config = config
self.rank = rank
self.logger = logger or logging.getLogger(__name__)
self.device = torch.device(
f"cuda:{rank}" if torch.cuda.is_available() else "cpu")
self.loss_fn = loss_fn or (lambda x, y: torch.mean((x - y.squeeze(-1))**2))
if config.use_wandb:
self.metrics_logger = WandBLogger(config.wandb_project, config.__dict__,
rank)
if config.distributed:
self.dist_manager = DistributedManager(
world_size=len(config.gpu_ids),
rank=rank,
master_addr=config.master_addr,
master_port=config.master_port,
)
self.dist_manager.setup()
self.model = self._setup_distributed_model()
def _setup_distributed_model(self) -> nn.Module:
"""Configure model for distributed training."""
self.model = self.model.to(self.device)
return DDP(self.model,
device_ids=[self.config.gpu_ids[self.rank]],
output_device=self.config.gpu_ids[self.rank])
def _create_dataloader(self, dataset: Dataset, is_train: bool) -> DataLoader:
"""Create appropriate DataLoader based on training configuration.
Args:
dataset: Dataset to create loader for.
is_train: Whether this is for training (affects shuffling).
Returns:
DataLoader instance.
"""
if self.config.distributed:
sampler = torch.utils.data.distributed.DistributedSampler(
dataset,
num_replicas=len(self.config.gpu_ids),
rank=dist.get_rank(),
shuffle=is_train)
else:
sampler = None
return DataLoader(
dataset,
batch_size=self.config.batch_size,
shuffle=(is_train and not self.config.distributed),
sampler=sampler,
)
def _quantile_loss(self, pred: torch.Tensor, actual: torch.Tensor,
quantile: float) -> torch.Tensor:
"""Calculates quantile loss.
Args:
pred: Predicted values
actual: Actual values
quantile: Quantile at which loss is computed
Returns:
Quantile loss
"""
dev = actual - pred
loss_first = dev * quantile
loss_second = -dev * (1.0 - quantile)
return 2 * torch.where(loss_first >= 0, loss_first, loss_second)
def _process_batch(self, batch: List[torch.Tensor]) -> tuple:
"""Process a single batch of data.
Args:
batch: List of input tensors.
Returns:
Tuple of (loss, predictions).
"""
x_context, x_padding, freq, x_future = [
t.to(self.device, non_blocking=True) for t in batch
]
predictions = self.model(x_context, x_padding.float(), freq)
predictions_mean = predictions[..., 0]
last_patch_pred = predictions_mean[:, -1, :]
loss = self.loss_fn(last_patch_pred, x_future.squeeze(-1))
if self.config.use_quantile_loss:
quantiles = self.config.quantiles or create_quantiles()
for i, quantile in enumerate(quantiles):
last_patch_quantile = predictions[:, -1, :, i + 1]
loss += torch.mean(
self._quantile_loss(last_patch_quantile, x_future.squeeze(-1),
quantile))
return loss, predictions
def _train_epoch(self, train_loader: DataLoader,
optimizer: torch.optim.Optimizer) -> float:
"""Train for one epoch in a distributed setting.
Args:
train_loader: DataLoader for training data.
optimizer: Optimizer instance.
Returns:
Average training loss for the epoch.
"""
self.model.train()
total_loss = 0.0
num_batches = len(train_loader)
for batch in train_loader:
loss, _ = self._process_batch(batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
total_loss += loss.item()
avg_loss = total_loss / num_batches
if self.config.distributed:
avg_loss_tensor = torch.tensor(avg_loss, device=self.device)
dist.all_reduce(avg_loss_tensor, op=dist.ReduceOp.SUM)
avg_loss = (avg_loss_tensor / dist.get_world_size()).item()
return avg_loss
def _validate(self, val_loader: DataLoader) -> float:
"""Perform validation.
Args:
val_loader: DataLoader for validation data.
Returns:
Average validation loss.
"""
self.model.eval()
total_loss = 0.0
num_batches = len(val_loader)
with torch.no_grad():
for batch in val_loader:
loss, _ = self._process_batch(batch)
total_loss += loss.item()
avg_loss = total_loss / num_batches
if self.config.distributed:
avg_loss_tensor = torch.tensor(avg_loss, device=self.device)
dist.all_reduce(avg_loss_tensor, op=dist.ReduceOp.SUM)
avg_loss = (avg_loss_tensor / dist.get_world_size()).item()
return avg_loss
def finetune(self, train_dataset: Dataset,
val_dataset: Dataset) -> Dict[str, Any]:
"""Train the model.
Args:
train_dataset: Training dataset.
val_dataset: Validation dataset.
Returns:
Dictionary containing training history.
"""
self.model = self.model.to(self.device)
train_loader = self._create_dataloader(train_dataset, is_train=True)
val_loader = self._create_dataloader(val_dataset, is_train=False)
optimizer = torch.optim.Adam(self.model.parameters(),
lr=self.config.learning_rate,
weight_decay=self.config.weight_decay)
history = {"train_loss": [], "val_loss": [], "learning_rate": []}
self.logger.info(
f"Starting training for {self.config.num_epochs} epochs...")
self.logger.info(f"Training samples: {len(train_dataset)}")
self.logger.info(f"Validation samples: {len(val_dataset)}")
try:
for epoch in range(self.config.num_epochs):
train_loss = self._train_epoch(train_loader, optimizer)
val_loss = self._validate(val_loader)
current_lr = optimizer.param_groups[0]["lr"]
metrics = {
"train_loss": train_loss,
"val_loss": val_loss,
"learning_rate": current_lr,
"epoch": epoch + 1,
}
if self.config.use_wandb:
self.metrics_logger.log_metrics(metrics)
history["train_loss"].append(train_loss)
history["val_loss"].append(val_loss)
history["learning_rate"].append(current_lr)
if self.rank == 0:
self.logger.info(
f"[Epoch {epoch+1}] Train Loss: {train_loss:.4f} | Val Loss: {val_loss:.4f}"
)
except KeyboardInterrupt:
self.logger.info("Training interrupted by user")
if self.config.distributed:
self.dist_manager.cleanup()
if self.config.use_wandb:
self.metrics_logger.close()
return {"history": history}
import apache_beam as beam
import logging
import torch
import numpy as np
import timesfm
from os import path
from timesfm import TimesFm, TimesFmCheckpoint, TimesFmHparams
from timesfm.pytorch_patched_decoder import PatchedTimeSeriesDecoder
from huggingface_hub import snapshot_download
from apache_beam.io.gcp.gcsio import GcsIO # Add this import
from torch.utils.data import Dataset
from google.cloud import storage
from typing import Tuple
class TimeSeriesDataset(Dataset):
"""Dataset for time series data compatible with TimesFM."""
def __init__(
self,
series: np.ndarray,
context_length: int,
horizon_length: int,
freq_type: int = 0):
"""
Initialize dataset.
Args:
series: Time series data
context_length: Number of past timesteps to use as input
horizon_length: Number of future timesteps to predict
freq_type: Frequency type (0, 1, or 2)
"""
if freq_type not in [0, 1, 2]:
raise ValueError("freq_type must be 0, 1, or 2")
self.series = series
self.context_length = context_length
self.horizon_length = horizon_length
self.freq_type = freq_type
self._prepare_samples()
def _prepare_samples(self) -> None:
"""Prepare sliding window samples from the time series."""
self.samples = []
total_length = self.context_length + self.horizon_length
for start_idx in range(0, len(self.series) - total_length + 1):
end_idx = start_idx + self.context_length
x_context = self.series[start_idx:end_idx]
x_future = self.series[end_idx:end_idx + self.horizon_length]
self.samples.append((x_context, x_future))
def __len__(self) -> int:
return len(self.samples)
def __getitem__(
self, index: int
) -> Tuple[torch.Tensor, torch.Tensor, torch.Tensor, torch.Tensor]:
x_context, x_future = self.samples[index]
x_context = torch.tensor(x_context, dtype=torch.float32)
x_future = torch.tensor(x_future, dtype=torch.float32)
input_padding = torch.zeros_like(x_context)
freq = torch.tensor([self.freq_type], dtype=torch.long)
return x_context, input_padding, freq, x_future
def prepare_datasets(
series: np.ndarray,
context_length: int,
horizon_length: int,
freq_type: int = 0,
train_split: float = 0.8) -> Tuple[Dataset, Dataset]:
"""
Prepare training and validation datasets from time series data.
Args:
series: Input time series data
context_length: Number of past timesteps to use
horizon_length: Number of future timesteps to predict
freq_type: Frequency type (0, 1, or 2)
train_split: Fraction of data to use for training
Returns:
Tuple of (train_dataset, val_dataset)
"""
train_size = int(len(series) * train_split)
train_data = series[:train_size]
val_data = series[train_size:]
# Create datasets with specified frequency type
train_dataset = TimeSeriesDataset(
train_data,
context_length=context_length,
horizon_length=horizon_length,
freq_type=freq_type)
val_dataset = TimeSeriesDataset(
val_data,
context_length=context_length,
horizon_length=horizon_length,
freq_type=freq_type)
return train_dataset, val_dataset
class BatchContinuousAndOrderedFn(beam.DoFn):
"""
A stateful DoFn that buffers elements, keeps them sorted, and emits
a batch only when a full, continuous sequence of points is available.
Includes detailed logging for debugging.
"""
BUFFER_STATE = ReadModifyWriteStateSpec('buffer', PickleCoder())
def __init__(self, batch_size, expected_interval_seconds=1):
self.batch_size = batch_size
self.interval = expected_interval_seconds
# NEW LOGGING: Counter to avoid logging on every single element
self.counter = 0
def process(self, element, buffer=beam.DoFn.StateParam(BUFFER_STATE)):
key, data = element
timestamp = data['timestamp']
value = data['value']
# Increment the counter
self.counter += 1
current_buffer = buffer.read() or []
current_buffer.append((timestamp, value))
current_buffer.sort(key=lambda x: x[0])
# NEW LOGGING: Periodically log the buffer status
if self.counter % 100 == 0 and current_buffer:
logging.info(
f"Batching buffer now contains {len(current_buffer)} points. "
f"Timestamps range from {current_buffer[0][0]} to {current_buffer[-1][0]}."
)
start_index = 0
while start_index + self.batch_size <= len(current_buffer):
is_continuous = True
# Check for continuity in the slice of the buffer we are considering
for i in range(start_index, start_index + self.batch_size - 1):
ts1_seconds = current_buffer[i][0].seconds()
ts2_seconds = current_buffer[i + 1][0].seconds()
if ts2_seconds - ts1_seconds != self.interval:
is_continuous = False
# If a gap is found, we should stop and wait for more data.
# We can't proceed past this point because the buffer is sorted.
logging.info(
f"Gap detected at index {i}. "
f"Timestamp {current_buffer[i][0]} is followed by {current_buffer[i+1][0]}. "
f"Actual interval: {ts2_seconds - ts1_seconds}s, Expected: {self.interval}s. "
f"Waiting for missing data."
)
break
if not is_continuous:
# Since the buffer is sorted, a gap at this point means we can't form any more continuous batches.
break
# If we are here, the batch from start_index is continuous.
logging.info(f"Continuous sequence found! Emitting batch of size {self.batch_size} starting at index {start_index}.")
batch_to_yield = current_buffer[start_index : start_index + self.batch_size]
formatted_batch = [{'timestamp': ts, 'value': val} for ts, val in batch_to_yield]
yield formatted_batch
# Move the start_index to the next position after the yielded batch
start_index += self.batch_size
# After the loop, remove all the yielded elements from the buffer.
if start_index > 0:
current_buffer = current_buffer[start_index:]
buffer.write(current_buffer)
class RunFinetuningFn(beam.DoFn):
"""
Takes a batch of data, loads the LATEST model, runs fine-tuning,
and uploads the new model to GCS.
"""
def __init__(
self,
initial_model_path, # Renamed from base_model_path
finetuned_model_bucket,
finetuned_model_prefix,
hparams,
config):
# This is now a fallback for the very first run
self.initial_model_path = initial_model_path
self.finetuned_model_bucket = finetuned_model_bucket
self.finetuned_model_prefix = finetuned_model_prefix
self.hparams = hparams
self.config = config
self._storage_client = None
def setup(self):
self._storage_client = storage.Client()
def _get_latest_model_from_gcs(self):
"""Directly queries GCS for the most recently created model checkpoint."""
try:
bucket = self._storage_client.get_bucket(self.finetuned_model_bucket)
blobs = list(bucket.list_blobs(prefix=self.finetuned_model_prefix))
# Filter for actual model files and exclude the initial model if present
model_blobs = [b for b in blobs if b.name.endswith(".pth") and "initial" not in b.name]
if not model_blobs:
return None
# Find the blob with the latest creation time
latest_blob = max(model_blobs, key=lambda b: b.time_created)
latest_model_path = f"gs://{self.finetuned_model_bucket}/{latest_blob.name}"
return latest_model_path
except Exception as e:
logging.error(f"Error querying GCS for the latest model: {e}")
return None
# Add the side input parameter to the process method
def process(self, batch_of_data):
logging.info(
f"Received a batch of {len(batch_of_data)} points for finetuning.")
# If a finetuned model exists, use it. Otherwise, use the initial base model.
latest_model_path = self._get_latest_model_from_gcs()
if latest_model_path:
model_to_load = latest_model_path
logging.info(f"Continuously finetuning from latest model: {model_to_load}")
else:
model_to_load = self.initial_model_path
logging.info(f"No finetuned model found. Starting from initial model: {model_to_load}")
# batch_of_data.sort(key=lambda x: x[1]['timestamp'])
time_series_values = np.array([d['value'] for d in batch_of_data],
dtype=np.float32)
train_dataset, val_dataset = prepare_datasets(
series=time_series_values,
context_length=self.hparams.context_len,
horizon_length=self.hparams.horizon_len,
freq_type=self.config.freq_type,
train_split=0.8
)
logging.info(f"Training dataset size: {train_dataset.series.tolist()}")
logging.info(f"Validation dataset size: {val_dataset.series.tolist()}")
# Load the model (base or latest finetuned)
# The updated get_model function can handle both GCS and Hugging Face paths
model = get_model(
model_path=model_to_load, # Use the path we just determined
hparams=self.hparams,
load_weights=True
)
# 4. Run fine-tuning (same as before)
finetuner = TimesFMFinetuner(model, self.config)
finetuner.finetune(train_dataset=train_dataset, val_dataset=val_dataset)
# 5. Save and upload the new model (same as before)
from datetime import datetime
timestamp_str = datetime.utcnow().strftime('%Y%m%d%H%M%S')
model_filename = f"timesfm_finetuned_{timestamp_str}.pth"
local_path = f"/tmp/{model_filename}"
torch.save(model.state_dict(), local_path)
bucket = self._storage_client.bucket(self.finetuned_model_bucket)
blob_path = f"{self.finetuned_model_prefix}/{model_filename}"
blob = bucket.blob(blob_path)
blob.upload_from_filename(local_path)
logging.info(
f"Successfully uploaded new model to gs://{self.finetuned_model_bucket}/{blob_path}"
)
yield blob_path
def get_model(model_path: str, hparams: TimesFmHparams, load_weights: bool = False):
"""
Loads a TimesFM model from either a Hugging Face repo ID or a GCS path.
The `load_weights` argument is kept for signature consistency but is
effectively always True, as TimesFm handles loading.
"""
checkpoint_config = {}
# Case 1: The model path is a GCS URI.
# We download it to a local file and tell TimesFmCheckpoint to load from that path.
if model_path.startswith("gs://"):
logging.info(f"Preparing to load model from GCS path: {model_path}")
local_temp_path = f"/tmp/{path.basename(model_path)}"
with GcsIO().open(model_path, 'rb') as f_in, open(local_temp_path, 'wb') as f_out:
f_out.write(f_in.read())
# The key for a local file is 'path'
checkpoint_config['path'] = local_temp_path
# Case 2: The model path is a Hugging Face repository ID.
else:
logging.info(f"Preparing to load model from Hugging Face repo: {model_path}")
# The key for a Hugging Face repo is 'huggingface_repo_id'
checkpoint_config['huggingface_repo_id'] = model_path
# Initialize the TimesFm object correctly with the dynamically created checkpoint config.
# This single call handles model configuration and weight loading.
tfm = TimesFm(
hparams=hparams,
checkpoint=TimesFmCheckpoint(**checkpoint_config)
)
logging.info("Model loaded successfully inside get_model.")
# The `TimesFm` object holds the configured model instance.
# The model returned here will be a PatchedTimeSeriesDecoder instance with weights loaded.
return tfm._model
Load Time Series Data
https://www.kaggle.com/datasets/julienjta/nyc-taxi-traffic/data
import pandas as pd
from google.colab import auth
auth.authenticate_user()
auth.authenticate_user()
# Define the path to your file in the GCS bucket
gcs_path = 'gs://apache-beam-samples/anomaly_detection/timesfm-dataset-example/nyc_taxi_timeseries.csv'
# Read the CSV directly from GCS into a DataFrame
# All the gspread code is replaced by this single line
df = pd.read_csv(gcs_path)
# --- The rest of your processing code remains the same ---
# Convert 'value' column to a numpy array of integers
values_array = pd.to_numeric(df['value'], errors='coerce').astype(int).to_numpy()
# Create the list of (timestamp, value) tuples
input_data = []
for i in range(len(values_array)):
input_data.append((Timestamp(i + 1), values_array[i])) # Assuming Timestamp comes from pandas
print("DataFrame loaded from GCS:")
print(df.head())
print("\nInput data created successfully (first 5 entries):")
print(input_data[:5])
DataFrame loaded from GCS: Unnamed: 0 timestamp value 0 0 2014-07-01 0:00:00 10844 1 1 2014-07-01 0:30:00 8127 2 2 2014-07-01 1:00:00 6210 3 3 2014-07-01 1:30:00 4656 4 4 2014-07-01 2:00:00 3820 Input data created successfully (first 5 entries): [(Timestamp(1), np.int64(10844)), (Timestamp(2), np.int64(8127)), (Timestamp(3), np.int64(6210)), (Timestamp(4), np.int64(4656)), (Timestamp(5), np.int64(3820))]
Beam Pipeline Setup
import apache_beam as beam
from apache_beam.options.pipeline_options import PipelineOptions
from apache_beam.pvalue import AsDict, AsSingleton
from apache_beam.transforms.periodicsequence import PeriodicImpulse
import logging
import os
import json
import timesfm
from apache_beam.utils.timestamp import Timestamp
import csv
from apache_beam.ml.inference.base import RunInference
from apache_beam.ml.inference.utils import WatchFilePattern
import typing
from google.colab import userdata
import apache_beam.transforms.window as window
logging.getLogger().setLevel(logging.INFO)
# --- Pipeline Configuration ---
PROJECT_ID = os.environ.get("GCP_PROJECT", "apache-beam-testing")
REGION = os.environ.get("GCP_REGION", "us-central1")
TEMP_LOCATION = "gs://apache-beam-testing-temp/timesfm_anomaly_detection/temp"
STAGING_LOCATION = "gs://apache-beam-testing-temp/timesfm_anomaly_detection/staging"
FINETUNED_MODEL_BUCKET = "apache-beam-testing-temp"
FINETUNED_MODEL_PREFIX = "timesfm_anomaly_detection/finetuned-models/timesfm/checkpoints"
# --- Model & Window Parameters ---
CONTEXT_LEN = 512
HORIZON_LEN = 128
WINDOW_SIZE = CONTEXT_LEN + HORIZON_LEN
SLIDE_INTERVAL = HORIZON_LEN
EXPECTED_INTERVAL = 1
INITIAL_MODEL = "google/timesfm-1.0-200m-pytorch"
MODEL_CHECK_INTERVAL_SECONDS = 10 # Check for a new model every 5 seconds
FINETUNING_BATCH_SIZE = 7680 # 9600 # make larger later. minimum is WINDOW_SIZE for validation and training
FINETUNE_CONFIG = FinetuningConfig(
batch_size=128,
num_epochs=5,
learning_rate=1e-4,
use_wandb=False,
freq_type=0, # should change based on your data
log_every_n_steps=10,
val_check_interval=0.5,
use_quantile_loss=True
)
#Change to Dataflow if needed
options = PipelineOptions([
"--streaming",
"--environment_type=LOOPBACK",
"--runner=PrismRunner",
"--logging_level=INFO",
"--job_server_timeout=3600"
])
# HParams for the model
hparams = timesfm.TimesFmHparams(
backend="gpu",
per_core_batch_size=32,
horizon_len=HORIZON_LEN,
context_len=CONTEXT_LEN,
)
model_handler = DynamicTimesFmModelHandler(model_uri=INITIAL_MODEL, hparams=hparams)
def print_and_pass_through(label):
def logger(element):
print(f"--- {label} --- \nELEMENT: %s", element)
return element
return logger
class CustomJsonEncoder(json.JSONEncoder):
"""A custom JSON encoder that knows how to handle Beam's Timestamp objects."""
def default(self, obj):
if isinstance(obj, Timestamp):
# Convert Timestamp to a standard, readable ISO 8601 string format
return obj.micros // 1e6
# For all other types, fall back to the default behavior
if isinstance(obj, np.integer):
return int(obj)
# 3. Handle NumPy float types (this will fix your float32 error)
if isinstance(obj, np.floating):
return float(obj)
# 4. Handle NumPy arrays
if isinstance(obj, np.ndarray):
return obj.tolist()
# For all other types, fall back to the default behavior
return super().default(obj)
return json.JSONEncoder.default(self, obj)
class WritePlotDataAndPassThrough(beam.DoFn):
"""
A DoFn that writes plotting data to a file as a side effect
and then passes the original, unmodified element downstream.
"""
def __init__(self, output_path):
self._output_path = output_path
self._file_handle = None
def setup(self):
self._file_handle = open(self._output_path, 'a')
def process(self, element):
_original_window, payload_dict = element
# ✅ FIX: Use the custom encoder to handle Timestamp objects
json_record = json.dumps(payload_dict, cls=CustomJsonEncoder)
self._file_handle.write(json_record + '\n')
# Pass the original element through, with the Timestamp object intact
yield element
def teardown(self):
if self._file_handle:
self._file_handle.close()
# =================================================================
# 1. Get Latest Model Path (Side Input) - WatchFilePattern is not
# currently supported on Prism. Uncomment the following to run
# on Dataflow
# =================================================================
# model_pattern = os.path.join(
# f"gs://{FINETUNED_MODEL_BUCKET}", FINETUNED_MODEL_PREFIX, "*.pth"
# )
# model_metadata_pcoll = (
# "WatchForNewModels" >> WatchFilePattern(
# file_pattern=model_pattern,
# interval=MODEL_CHECK_INTERVAL_SECONDS
# )
# | "PrintModelLocation" >> beam.Map(print_and_pass_through("Model Location"))
# )
# =================================================================
# Ingest and Window Raw Data
# =================================================================
windowed_data = (
PeriodicImpulse(data=input_data, fire_interval=0.01)
| "AddKey" >> beam.WithKeys(lambda x: 0)
| "ApplySlidingWindow" >> beam.ParDo(
OrderedSlidingWindowFn(window_size=WINDOW_SIZE, slide_interval=SLIDE_INTERVAL))
| "FillGaps" >> beam.ParDo(FillGapsFn(expected_interval=EXPECTED_INTERVAL)).with_output_types(
typing.Tuple[int, typing.Tuple[Timestamp, Timestamp, typing.List[float]]])
| "Skip NaN Values for now" >> beam.Filter(
lambda batch: 'NaN' not in batch[1][2])
| "PrintWindowedData" >> beam.Map(print_and_pass_through("Windowed Data"))
)
# =================================================================
# Detect Anomalies using the Latest Model
# =================================================================
inference_results = (
"DetectAnomalies" >> RunInference(
model_handler=model_handler,
# model_metadata_pcoll=model_metadata_pcoll
)
| "PrintInference" >> beam.Map(print_and_pass_through("Inference Results"))
)
# NEW BRANCH: For plotting. It takes the payload dictionary, converts
# it to JSON, and writes it to a file.
plotting_data_output = (
"WritePlotDataAsSideEffect" >> beam.ParDo(
WritePlotDataAndPassThrough('plot_data_original.jsonl'))
)
def format_for_llm(result_tuple):
"""
Takes the output of RunInference (a PredictionResult) and formats it
into the dictionary structure needed by the LLMClassifierFn.
"""
original_window_data, result_dict = result_tuple
list_of_anomalies = result_dict['anomalies']
key, (window_start_ts, _, values_array) = original_window_data
return (key, {
'key': key,
'window_start_ts': window_start_ts,
'values_array': values_array,
'anomalies': list_of_anomalies if list_of_anomalies else []
})
data_for_llm = (
"FormatForLLM" >> beam.Map(format_for_llm)
| "PrintDataForLLM" >> beam.Map(print_and_pass_through("Data for LLM"))
)
# =================================================================
# Classify with LLM and Create Clean Data for Finetuning
# =================================================================
api_key = "AIzaSyCB_g6tq3eBFtB3BsshdGotLkUkTsCyApY" #userdata.get('GEMINI_API_KEY')
llm_classifier = (
"LLMClassifierAndImputer" >> beam.ParDo(
LLMClassifierFn(
secret=api_key,
slide_interval=SLIDE_INTERVAL,
expected_interval_secs=EXPECTED_INTERVAL
)
)
# | "PrintLLMResults" >> beam.Map(print_and_pass_through("LLM Results"))
)
# # =================================================================
# # Batch Clean Data and Trigger Finetuning
# # =================================================================
finetuning_job_input = (
"KeyForBatching" >> beam.WithKeys(lambda _: "finetune_batch")
# | "BatchAndTrigger" >> beam.ParDo(BatchAndTriggerFinetuningFn(FINETUNING_BATCH_SIZE))
| "BatchAndTrigger" >> beam.ParDo(
BatchContinuousAndOrderedFn(
FINETUNING_BATCH_SIZE,
expected_interval_seconds=1
)
)
| "PrintFinetuningJobInput" >> beam.Map(print_and_pass_through("Finetuning Job Input"))
)
# # =================================================================
# # Run Finetuning and Save New Model to GCS
# # =================================================================
finetuning = (
"RunFinetuning" >> beam.ParDo(
RunFinetuningFn(
initial_model_path="google/timesfm-1.0-200m-pytorch",
finetuned_model_bucket=FINETUNED_MODEL_BUCKET,
finetuned_model_prefix=FINETUNED_MODEL_PREFIX,
hparams=hparams,
config=FINETUNE_CONFIG
),
)
)
WARNING:apache_beam.transforms.core:('No iterator is returned by the process method in %s.', <class '__main__.LLMClassifierFn'>)
Beam Pipeline
with beam.Pipeline(options=options) as p:
(p
| windowed_data
| inference_results
| plotting_data_output # comment this line if you dont want to save plot data
| data_for_llm
| llm_classifier
| finetuning_job_input
| finetuning
)
Plot Data (Original)
import json
import numpy as np
import matplotlib.pyplot as plt
CONTEXT_LEN = 512
HORIZON_LEN = 128
def plot_anomalies_and_forecast(
values_array,
all_anomalies,
all_predicted_values,
all_q20_values,
all_q30_values,
all_q70_values,
all_q80_values,
title_suffix="",
x_lims=None,
min_outlier_score_for_plot=0,
context_len=512,
output_filename="plot.png"
):
print(len(all_anomalies))
# The key from your file is 'outlier_score'
filtered_anomalies = [a for a in all_anomalies if a['outlier_score'] >= min_outlier_score_for_plot]
# The key from your file is 'timestamp'
anomaly_indices = [(a['timestamp'] - HORIZON_LEN) for a in filtered_anomalies]
anomaly_values = [a['actual_value'] for a in filtered_anomalies]
Q1 = np.nanmean([all_q20_values, all_q30_values], axis=0)
Q3 = np.nanmean([all_q70_values, all_q80_values], axis=0)
IQR = Q3 - Q1
upper_thresh = Q3 + 1.5 * IQR
lower_thresh = Q1 - 1.5 * IQR
plt.figure(figsize=(18, 9))
# This now plots the correct original data for the horizon
plt.plot(values_array[context_len:], label='Original Time Series', color='blue', alpha=0.7, linewidth=1.5)
plt.plot(all_predicted_values, label='Predicted Mean', color='green', linestyle='--', linewidth=1.5)
plt.plot(lower_thresh, label='Lower Threshold', color='orange', linestyle=':', linewidth=1.2)
plt.plot(upper_thresh, label='Upper Threshold', color='purple', linestyle=':', linewidth=1.2)
plt.scatter([i - context_len for i in anomaly_indices], anomaly_values,
color='red', s=70, zorder=5,
label=f'Detected Anomalies (Score >= {min_outlier_score_for_plot:.1f})',
marker='o', edgecolors='black', linewidths=0.8)
plt.title(f'Time Series Anomaly Detection {title_suffix}')
plt.xlabel('Time Index')
plt.ylabel('Value')
if x_lims:
plt.xlim(x_lims[0], x_lims[1])
plt.legend()
plt.grid(True, linestyle='--', alpha=0.6)
plt.tight_layout()
# plt.savefig(output_filename) # Save the plot to a file
plt.show()
plt.close() # Close the figure to free memory
# --- Main Script Logic ---
# 1. Read and parse the data from the Beam output file
all_window_data = []
# Make sure 'plot_data.jsonl' is in the same directory as this script
try:
with open('plot_data_original.jsonl', 'r') as f:
for line in f:
# Check for empty lines that might have been added
if line.strip():
all_window_data.append(json.loads(line))
except FileNotFoundError:
print("Error: 'plot_data.jsonl' not found. Please make sure the file is in the correct directory.")
exit()
# 2. Sort data by timestamp to ensure the correct order
all_window_data.sort(key=lambda x: x['start_ts_micros'])
# 3. Reconstruct the full data arrays
all_anomalies = []
all_predicted_values = []
all_q20_values = []
all_q30_values = []
all_q70_values = []
all_q80_values = []
all_actual_horizon_values = [] # This will hold the real "blue line" data
for window_data in all_window_data:
all_predicted_values.extend(window_data['predicted_values'])
all_q20_values.extend(window_data['q20_values'])
all_q30_values.extend(window_data['q30_values'])
all_q70_values.extend(window_data['q70_values'])
all_q80_values.extend(window_data['q80_values'])
# Populate the list with the actual values from the file
all_actual_horizon_values.extend(window_data.get('actual_horizon_values', []))
all_anomalies.extend(window_data.get('anomalies', []))
# 4. Convert lists to NumPy arrays
all_predicted_values = np.array(all_predicted_values)
all_q20_values = np.array(all_q20_values)
all_q30_values = np.array(all_q30_values)
all_q70_values = np.array(all_q70_values)
all_q80_values = np.array(all_q80_values)
# 5. Construct the `values_array` using the REAL data from your file
context_len = 512
# Create a dummy context so the array has the right shape for the plotting function.
# The first real value is used to make the context visually seamless.
if all_actual_horizon_values:
dummy_context = [all_actual_horizon_values[0]] * context_len
values_array = np.array(dummy_context + all_actual_horizon_values)
else:
# Fallback in case the file is empty or missing the actual_horizon_values key
print("Warning: 'actual_horizon_values' not found. The original time series plot will be empty.")
total_len = context_len + len(all_predicted_values)
values_array = np.zeros(total_len)
# 6. Call the plotting functions
if values_array.any():
# Plotting function for full graph
plot_anomalies_and_forecast(
values_array, all_anomalies, all_predicted_values,
all_q20_values, all_q30_values, all_q70_values, all_q80_values,
title_suffix="(Full Graph with Correct Data)",
min_outlier_score_for_plot=1, # Set a score threshold
context_len=context_len,
output_filename="full_graph_correct.png"
)
# Plotting function for zoomed-in graphs - feel free to change
zoom_ranges = [(2000, 2500), (8300, 9000), (9000, 9600)]
for i, (start_idx, end_idx) in enumerate(zoom_ranges):
# Adjust x_lims for the fact that the plotted array is sliced by context_len
plot_x_start = max(0, start_idx)
plot_x_end = end_idx
plot_anomalies_and_forecast(
values_array, all_anomalies, all_predicted_values,
all_q20_values, all_q30_values, all_q70_values, all_q80_values,
title_suffix=f"(Zoomed In: {start_idx} to {end_idx})",
x_lims=(plot_x_start, plot_x_end),
min_outlier_score_for_plot=1,
context_len=context_len,
output_filename=f"zoomed_graph_correct_{i}.png"
)
print("Plots have been generated and saved with the corrected original data.")
else:
print("No data found to plot.")
948
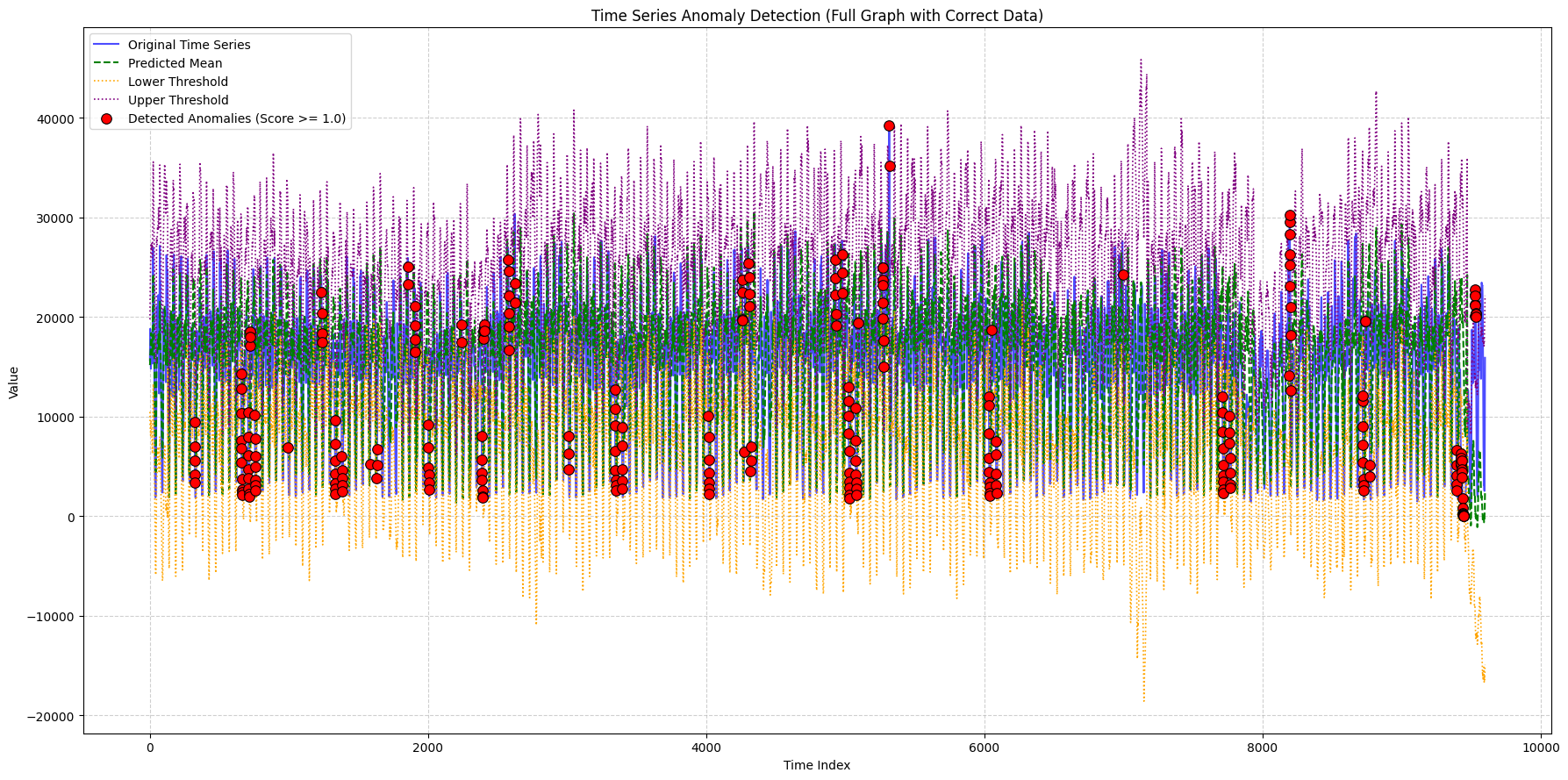
948
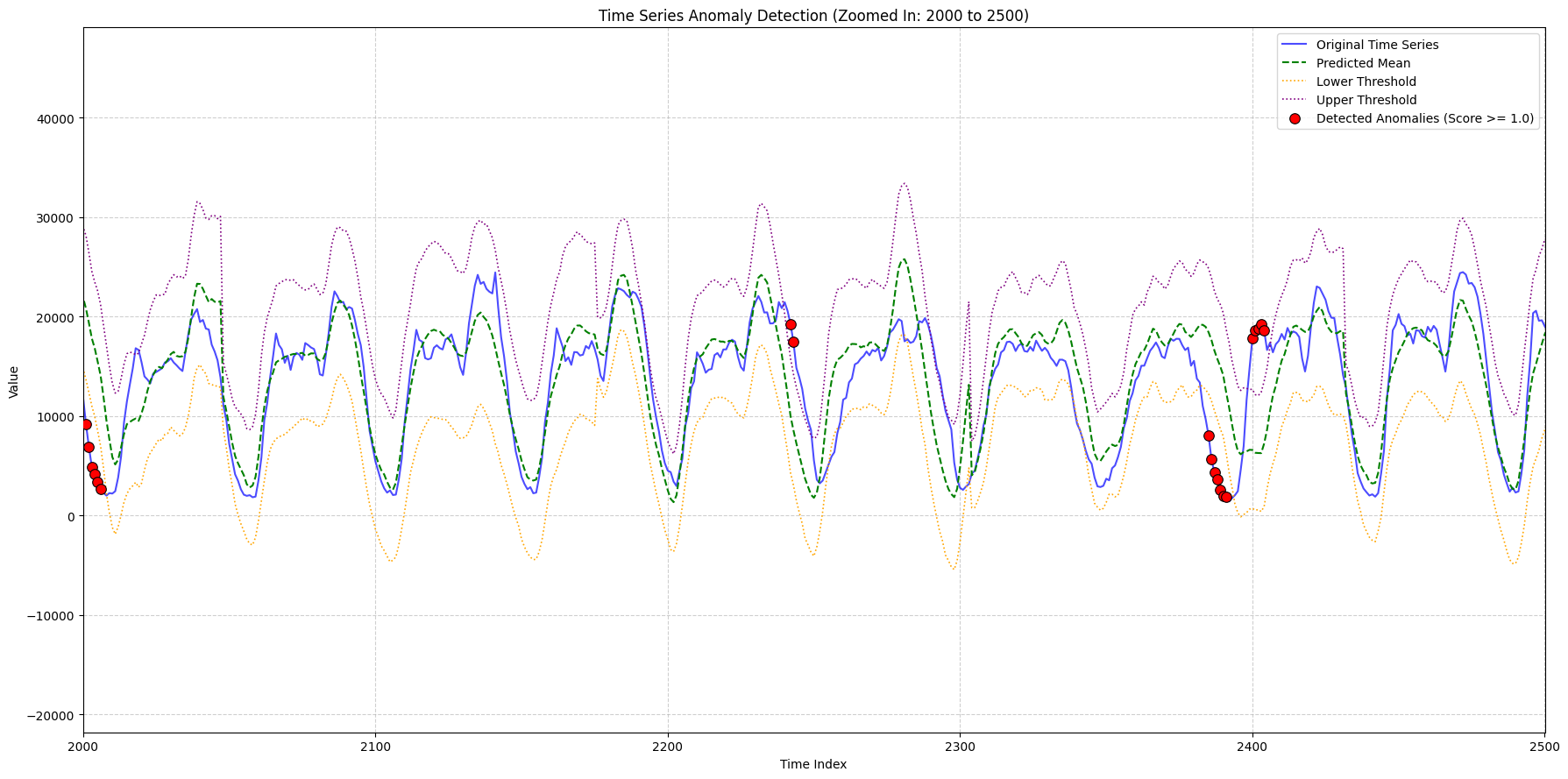
948
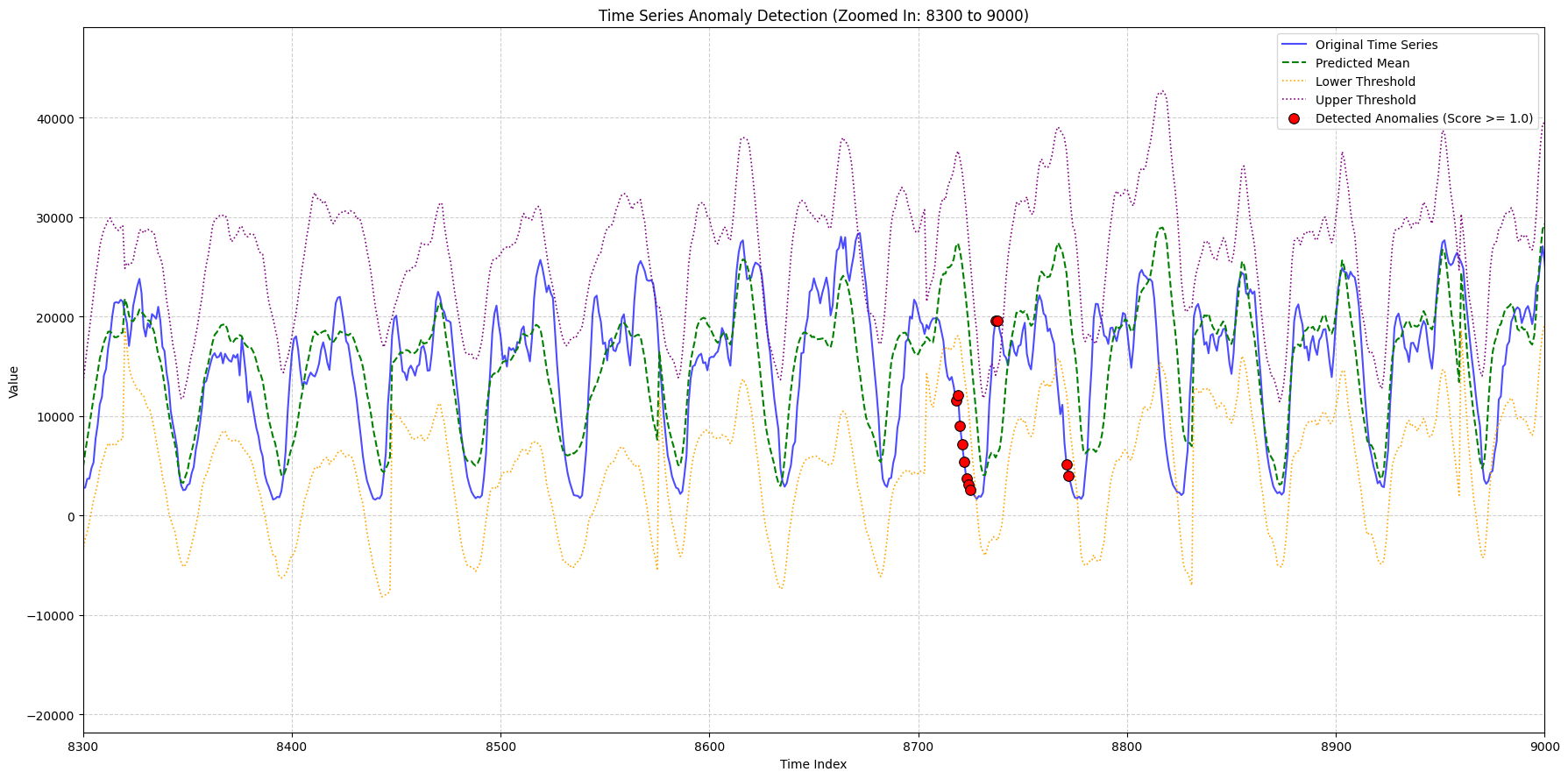
948
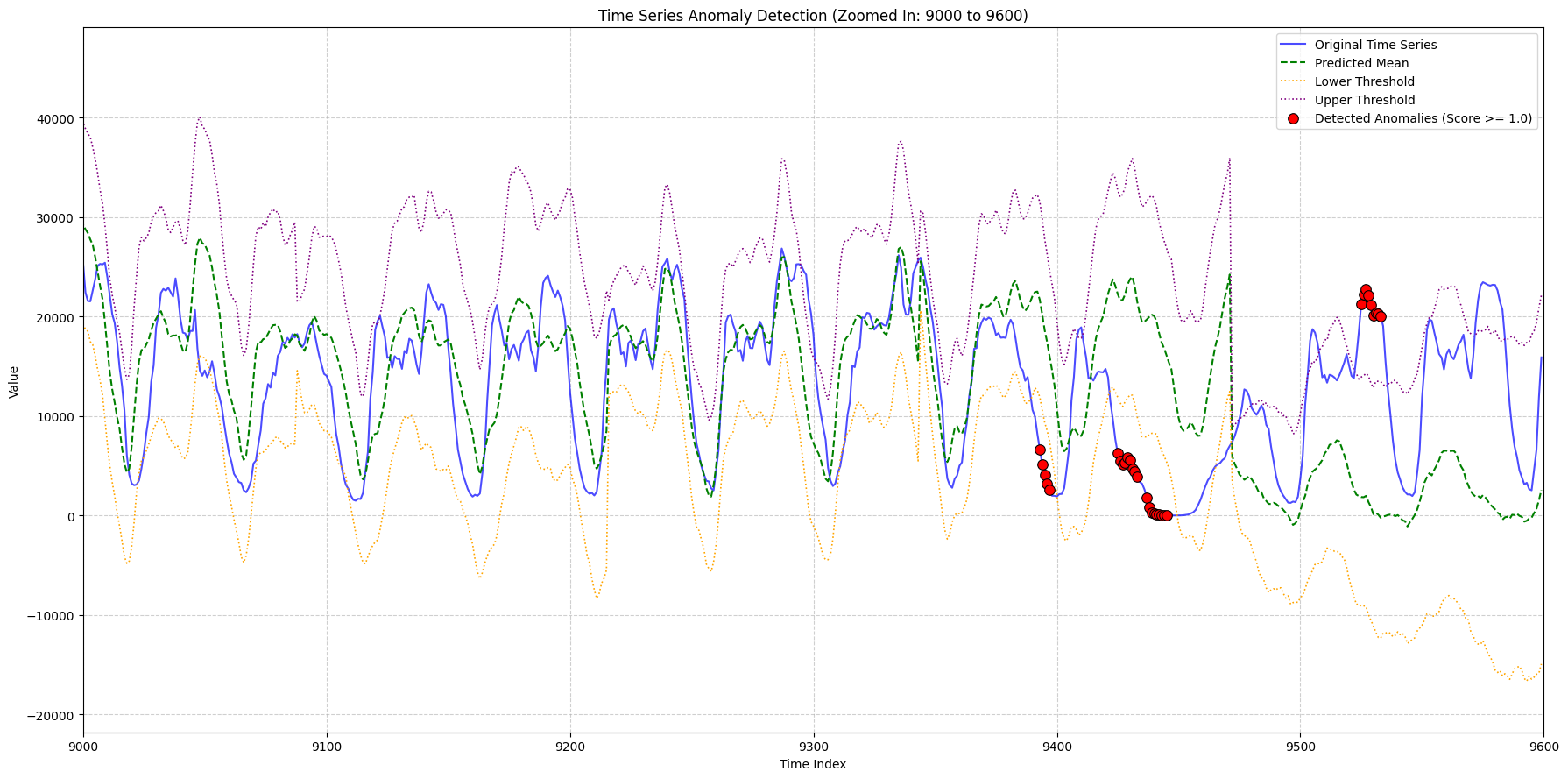
Plots have been generated and saved with the corrected original data.
Finetuned Model Predictions
import apache_beam as beam
from apache_beam.options.pipeline_options import PipelineOptions
from apache_beam.pvalue import AsDict, AsSingleton
from apache_beam.transforms.periodicsequence import PeriodicImpulse
import logging
import os
import json
import timesfm
from apache_beam.utils.timestamp import Timestamp
import csv
from apache_beam.ml.inference.base import RunInference
from apache_beam.ml.inference.utils import WatchFilePattern
import typing
from google.colab import userdata
import apache_beam.transforms.window as window
logging.getLogger().setLevel(logging.INFO)
# --- Pipeline Configuration ---
PROJECT_ID = os.environ.get("GCP_PROJECT", "apache-beam-testing")
REGION = os.environ.get("GCP_REGION", "us-central1")
TEMP_LOCATION = "gs://apache-beam-testing-temp/timesfm_anomaly_detection/temp"
STAGING_LOCATION = "gs://apache-beam-testing-temp/timesfm_anomaly_detection/staging"
FINETUNED_MODEL_BUCKET = "apache-beam-testing-temp"
FINETUNED_MODEL_PREFIX = "timesfm_anomaly_detection/finetuned-models/timesfm/checkpoints"
# --- Model & Window Parameters ---
CONTEXT_LEN = 512
HORIZON_LEN = 128
WINDOW_SIZE = CONTEXT_LEN + HORIZON_LEN
SLIDE_INTERVAL = HORIZON_LEN
EXPECTED_INTERVAL = 1
# go to checkpoints bucket and select the correct model path
INITIAL_MODEL = "gs://apache-beam-testing-temp/timesfm_anomaly_detection/finetuned-models/timesfm/checkpoints/timesfm_finetuned_20250814192006.pth"
MODEL_CHECK_INTERVAL_SECONDS = 10 # Check for a new model every 5 seconds
FINETUNING_BATCH_SIZE = 7680 # 9600 # make larger later. minimum is WINDOW_SIZE for validation and training
FINETUNE_CONFIG = FinetuningConfig(
batch_size=128,
num_epochs=5,
learning_rate=1e-4,
use_wandb=False,
freq_type=0, # should change based on your data
log_every_n_steps=10,
val_check_interval=0.5,
use_quantile_loss=True
)
options = PipelineOptions([
"--streaming",
"--environment_type=LOOPBACK",
"--runner=PrismRunner",
"--logging_level=INFO",
"--job_server_timeout=3600"
])
# HParams for the model
hparams = timesfm.TimesFmHparams(
backend="gpu",
per_core_batch_size=32,
horizon_len=HORIZON_LEN,
context_len=CONTEXT_LEN,
)
model_handler = DynamicTimesFmModelHandler(model_uri=INITIAL_MODEL, hparams=hparams)
def print_and_pass_through(label):
def logger(element):
print(f"--- {label} --- \nELEMENT: %s", element)
return element
return logger
class CustomJsonEncoder(json.JSONEncoder):
"""A custom JSON encoder that knows how to handle Beam's Timestamp objects."""
def default(self, obj):
if isinstance(obj, Timestamp):
# Convert Timestamp to a standard, readable ISO 8601 string format
return obj.micros // 1e6
# For all other types, fall back to the default behavior
if isinstance(obj, np.integer):
return int(obj)
# 3. Handle NumPy float types (this will fix your float32 error)
if isinstance(obj, np.floating):
return float(obj)
# 4. Handle NumPy arrays
if isinstance(obj, np.ndarray):
return obj.tolist()
# For all other types, fall back to the default behavior
return super().default(obj)
return json.JSONEncoder.default(self, obj)
class WritePlotDataAndPassThrough(beam.DoFn):
"""
A DoFn that writes plotting data to a file as a side effect
and then passes the original, unmodified element downstream.
"""
def __init__(self, output_path):
self._output_path = output_path
self._file_handle = None
def setup(self):
self._file_handle = open(self._output_path, 'a')
def process(self, element):
_original_window, payload_dict = element
# ✅ FIX: Use the custom encoder to handle Timestamp objects
json_record = json.dumps(payload_dict, cls=CustomJsonEncoder)
self._file_handle.write(json_record + '\n')
# Pass the original element through, with the Timestamp object intact
yield element
def teardown(self):
if self._file_handle:
self._file_handle.close()
# =================================================================
# 1. Get Latest Model Path (Side Input) - WatchFilePattern is not
# currently supported on Prism. Uncomment the following to run
# on Dataflow
# =================================================================
# model_pattern = os.path.join(
# f"gs://{FINETUNED_MODEL_BUCKET}", FINETUNED_MODEL_PREFIX, "*.pth"
# )
# model_metadata_pcoll = (
# "WatchForNewModels" >> WatchFilePattern(
# file_pattern=model_pattern,
# interval=MODEL_CHECK_INTERVAL_SECONDS
# )
# | "PrintModelLocation" >> beam.Map(print_and_pass_through("Model Location"))
# )
# =================================================================
# Ingest and Window Raw Data
# =================================================================
windowed_data = (
PeriodicImpulse(data=input_data, fire_interval=0.01)
| "AddKey" >> beam.WithKeys(lambda x: 0)
| "ApplySlidingWindow" >> beam.ParDo(
OrderedSlidingWindowFn(window_size=WINDOW_SIZE, slide_interval=SLIDE_INTERVAL))
| "FillGaps" >> beam.ParDo(FillGapsFn(expected_interval=EXPECTED_INTERVAL)).with_output_types(
typing.Tuple[int, typing.Tuple[Timestamp, Timestamp, typing.List[float]]])
| "Skip NaN Values for now" >> beam.Filter(
lambda batch: 'NaN' not in batch[1][2])
| "PrintWindowedData" >> beam.Map(print_and_pass_through("Windowed Data"))
)
# =================================================================
# Detect Anomalies using the Latest Model
# =================================================================
inference_results = (
"DetectAnomalies" >> RunInference(
model_handler=model_handler,
# model_metadata_pcoll=model_metadata_pcoll
)
| "PrintInference" >> beam.Map(print_and_pass_through("Inference Results"))
)
# NEW BRANCH: For plotting. It takes the payload dictionary, converts
# it to JSON, and writes it to a file.
plotting_data_output = (
"WritePlotDataAsSideEffect" >> beam.ParDo(
WritePlotDataAndPassThrough('plot_data_finetuned.jsonl'))
)
with beam.Pipeline(options=options) as p:
(p
| windowed_data
| inference_results
| plotting_data_output
)
Plot Data (After Finetuning)
import json
import numpy as np
import matplotlib.pyplot as plt
def plot_anomalies_and_forecast(
values_array,
all_anomalies,
all_predicted_values,
all_q20_values,
all_q30_values,
all_q70_values,
all_q80_values,
title_suffix="",
x_lims=None,
min_outlier_score_for_plot=0,
context_len=512,
output_filename="plot.png"
):
# The key from your file is 'outlier_score'
filtered_anomalies = [a for a in all_anomalies if a['outlier_score'] >= min_outlier_score_for_plot]
# The key from your file is 'timestamp'
anomaly_indices = [(a['timestamp'] - 128) for a in filtered_anomalies]
anomaly_values = [a['actual_value'] for a in filtered_anomalies]
Q1 = np.nanmean([all_q20_values, all_q30_values], axis=0)
Q3 = np.nanmean([all_q70_values, all_q80_values], axis=0)
IQR = Q3 - Q1
upper_thresh = Q3 + 1.5 * IQR
lower_thresh = Q1 - 1.5 * IQR
plt.figure(figsize=(18, 9))
# This now plots the correct original data for the horizon
plt.plot(values_array[context_len:], label='Original Time Series', color='blue', alpha=0.7, linewidth=1.5)
plt.plot(all_predicted_values, label='Predicted Mean', color='green', linestyle='--', linewidth=1.5)
plt.plot(lower_thresh, label='Lower Threshold', color='orange', linestyle=':', linewidth=1.2)
plt.plot(upper_thresh, label='Upper Threshold', color='purple', linestyle=':', linewidth=1.2)
plt.scatter([i - context_len for i in anomaly_indices], anomaly_values,
color='red', s=70, zorder=5,
label=f'Detected Anomalies (Score >= {min_outlier_score_for_plot:.1f})',
marker='o', edgecolors='black', linewidths=0.8)
plt.title(f'Time Series Anomaly Detection {title_suffix}')
plt.xlabel('Time Index')
plt.ylabel('Value')
if x_lims:
plt.xlim(x_lims[0], x_lims[1])
plt.legend()
plt.grid(True, linestyle='--', alpha=0.6)
plt.tight_layout()
# plt.savefig(output_filename) # Save the plot to a file
plt.show()
plt.close() # Close the figure to free memory
# --- Main Script Logic ---
# 1. Read and parse the data from the Beam output file
all_window_data = []
# Make sure 'plot_data.jsonl' is in the same directory as this script
try:
with open('plot_data_finetuned.jsonl', 'r') as f:
for line in f:
# Check for empty lines that might have been added
if line.strip():
all_window_data.append(json.loads(line))
except FileNotFoundError:
print("Error: 'plot_data_finetuned.jsonl' not found. Please make sure the file is in the correct directory.")
exit()
# 2. Sort data by timestamp to ensure the correct order
all_window_data.sort(key=lambda x: x['start_ts_micros'])
# 3. Reconstruct the full data arrays
all_anomalies = []
all_predicted_values = []
all_q20_values = []
all_q30_values = []
all_q70_values = []
all_q80_values = []
all_actual_horizon_values = [] # This will hold the real "blue line" data
for window_data in all_window_data:
all_predicted_values.extend(window_data['predicted_values'])
all_q20_values.extend(window_data['q20_values'])
all_q30_values.extend(window_data['q30_values'])
all_q70_values.extend(window_data['q70_values'])
all_q80_values.extend(window_data['q80_values'])
# Populate the list with the actual values from the file
all_actual_horizon_values.extend(window_data.get('actual_horizon_values', []))
all_anomalies.extend(window_data.get('anomalies', []))
# 4. Convert lists to NumPy arrays
all_predicted_values = np.array(all_predicted_values)
print(len(all_predicted_values))
all_q20_values = np.array(all_q20_values)
all_q30_values = np.array(all_q30_values)
all_q70_values = np.array(all_q70_values)
all_q80_values = np.array(all_q80_values)
# 5. Construct the `values_array` using the REAL data from your file
context_len = 512
# Create a dummy context so the array has the right shape for the plotting function.
# The first real value is used to make the context visually seamless.
if all_actual_horizon_values:
dummy_context = [all_actual_horizon_values[0]] * context_len
values_array = np.array(dummy_context + all_actual_horizon_values)
else:
# Fallback in case the file is empty or missing the actual_horizon_values key
print("Warning: 'actual_horizon_values' not found. The original time series plot will be empty.")
total_len = context_len + len(all_predicted_values)
values_array = np.zeros(total_len)
# 6. Call the plotting functions
if values_array.any():
# Plotting function for full graph
plot_anomalies_and_forecast(
values_array, all_anomalies, all_predicted_values,
all_q20_values, all_q30_values, all_q70_values, all_q80_values,
title_suffix="(Full Graph with Correct Data)",
min_outlier_score_for_plot=5, # Set a score threshold
context_len=context_len,
output_filename="full_graph_correct.png"
)
# Plotting function for zoomed-in graphs - feel free to change
zoom_ranges = [(2000, 2500), (8300, 9000), (9000, 9600)]
for i, (start_idx, end_idx) in enumerate(zoom_ranges):
# Adjust x_lims for the fact that the plotted array is sliced by context_len
plot_x_start = max(0, start_idx)
plot_x_end = end_idx
plot_anomalies_and_forecast(
values_array, all_anomalies, all_predicted_values,
all_q20_values, all_q30_values, all_q70_values, all_q80_values,
title_suffix=f"(Zoomed In: {start_idx} to {end_idx})",
x_lims=(plot_x_start, plot_x_end),
min_outlier_score_for_plot=5,
context_len=context_len,
output_filename=f"zoomed_graph_correct_{i}.png"
)
print("Plots have been generated and saved with the corrected original data.")
else:
print("No data found to plot.")
9600
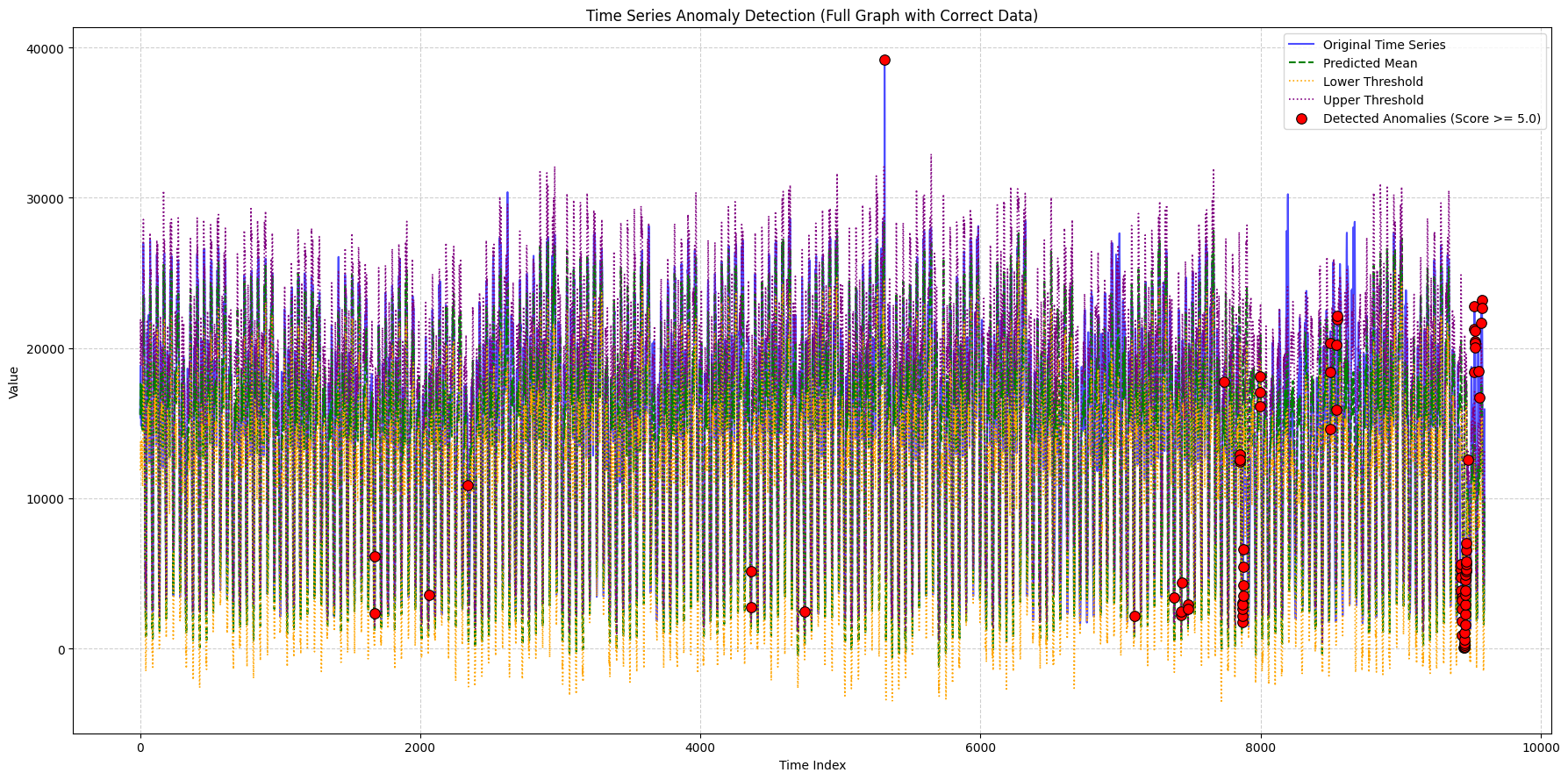
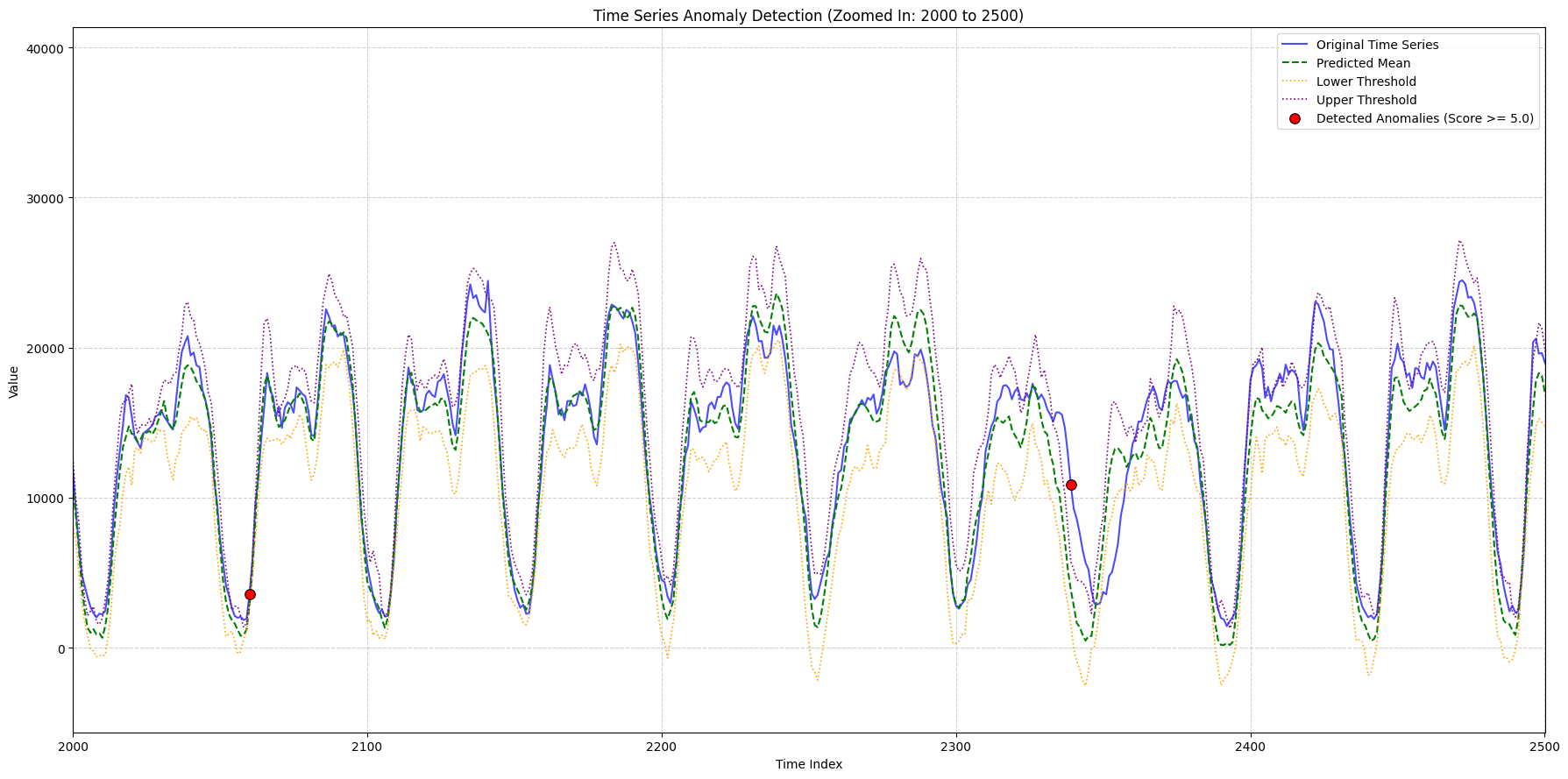
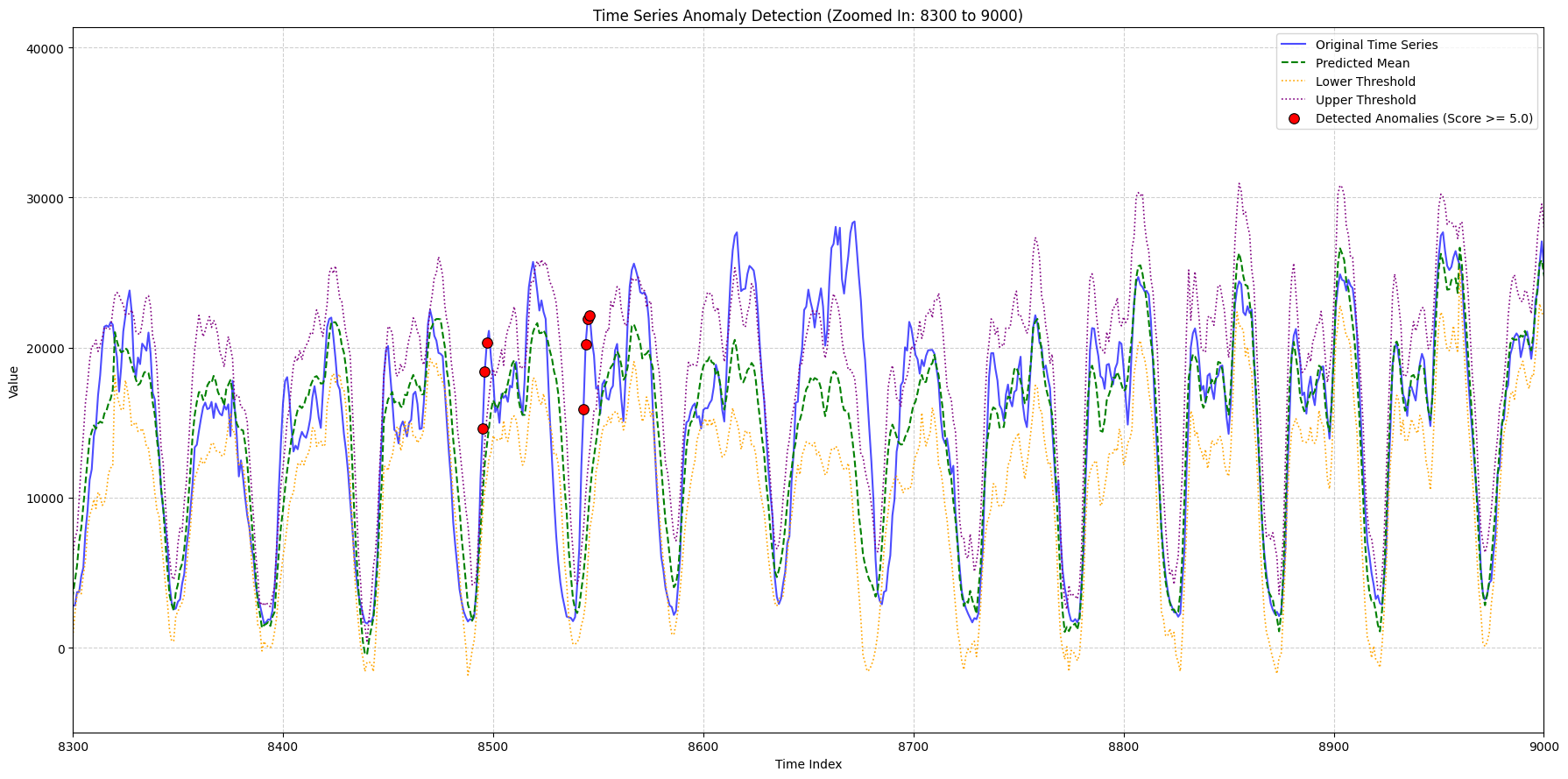
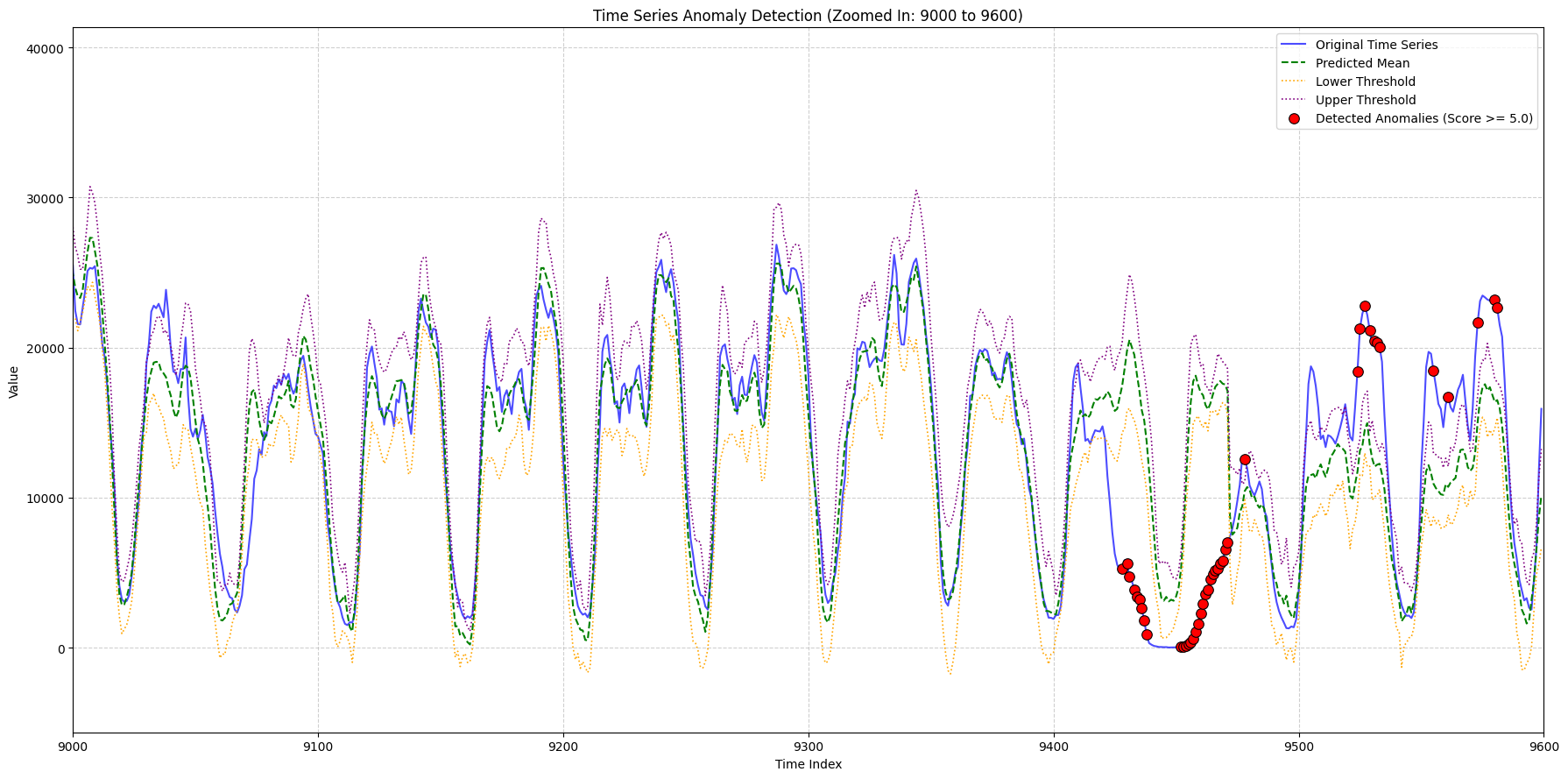
Plots have been generated and saved with the corrected original data.
Compare with RMSE (Original vs Finetuned on Test Data)
import json
import numpy as np
import pandas as pd
def calculate_model_rmse(file_path, finetuning_cutoff_ts, outlier_threshold):
"""
Loads model output data, reconstructs the time series, and calculates RMSE
on a test set after filtering outliers.
Args:
file_path (str): Path to the model's prediction data in JSONL format.
finetuning_cutoff_ts (int): Timestamp to split training and test data.
outlier_threshold (float): Outlier score at or above which to exclude points.
Returns:
float: The calculated Root Mean Squared Error.
"""
all_window_data = []
with open(file_path, 'r') as f:
for line in f:
if line.strip():
all_window_data.append(json.loads(line))
all_window_data.sort(key=lambda x: x['start_ts_micros'])
# --- Reconstruct the full time series from the windows ---
timestamps = []
all_predicted_values = []
all_actual_values = []
all_anomalies = []
current_ts = -1
if all_window_data:
# Initialize with the first window's start time
current_ts = all_window_data[0]['start_ts_micros'] // 1000000
for window_data in all_window_data:
# Extend the series lists
all_predicted_values.extend(window_data['predicted_values'])
all_actual_values.extend(window_data.get('actual_horizon_values', []))
all_anomalies.extend(window_data.get('anomalies', []))
# Reconstruct the timestamps for each predicted point
start_ts = window_data['start_ts_micros'] // 1000000
for _ in window_data['predicted_values']:
timestamps.append(start_ts)
start_ts += 1
# Create a lookup for outlier scores
outlier_scores_map = {item['timestamp']: item['outlier_score'] for item in all_anomalies}
# Ensure the actual values and predicted values align
min_len = min(len(timestamps), len(all_predicted_values), len(all_actual_values))
# --- Create a DataFrame for easy filtering and calculation ---
df = pd.DataFrame({
'timestamp': timestamps[:min_len],
'actual': all_actual_values[:min_len],
'predicted': all_predicted_values[:min_len]
})
df['outlier_score'] = df['timestamp'].map(outlier_scores_map).fillna(0.0)
# 1. Isolate the test set
df_test = df[df['timestamp'] > finetuning_cutoff_ts].copy()
print(f"\n--- Analyzing: {file_path} ---")
print(f"Test set size (before filtering): {len(df_test)} points")
# 2. Filter out anomalies based on the threshold
df_filtered = df_test[df_test['outlier_score'] < outlier_threshold]
num_outliers = len(df_test) - len(df_filtered)
print(f"Test set size (after filtering): {len(df_filtered)} points")
print(f"Removed {num_outliers} points with outlier_score >= {outlier_threshold}")
# 3. Calculate RMSE
y_true = df_filtered['actual']
y_pred = df_filtered['predicted']
rmse = np.sqrt(np.mean((y_true - y_pred)**2))
return rmse
# --- Configuration ---
FINETUNING_CUTOFF_TS = 8320
ORIGINAL_OUTLIER_THRESHOLD = 1.0
FINETUNED_OUTLIER_THRESHOLD = 5.0
# --- Execution & Comparison ---
original_rmse = calculate_model_rmse(
file_path="plot_data_original.jsonl",
finetuning_cutoff_ts=FINETUNING_CUTOFF_TS,
outlier_threshold=ORIGINAL_OUTLIER_THRESHOLD
)
finetuned_rmse = calculate_model_rmse(
file_path="plot_data_finetuned.jsonl",
finetuning_cutoff_ts=FINETUNING_CUTOFF_TS,
outlier_threshold=FINETUNED_OUTLIER_THRESHOLD
)
print("\n--- Final Results ---")
print(f"Original Model RMSE: {original_rmse:.2f}")
print(f"Fine-tuned Model RMSE: {finetuned_rmse:.2f}")
--- Analyzing: plot_data_original.jsonl --- Test set size (before filtering): 1407 points Test set size (after filtering): 1369 points Removed 38 points with outlier_score >= 1.0 --- Analyzing: plot_data_finetuned.jsonl --- Test set size (before filtering): 1407 points Test set size (after filtering): 1384 points Removed 23 points with outlier_score >= 5.0 --- Final Results --- Original Model RMSE: 7164.17 Fine-tuned Model RMSE: 3948.44

